The microtubule plus-end binding protein EB1 functions in root responses to touch and gravity signals in Arabidopsis
- PMID: 18281505
- PMCID: PMC2276450
- DOI: 10.1105/tpc.107.056846
The microtubule plus-end binding protein EB1 functions in root responses to touch and gravity signals in Arabidopsis
Abstract
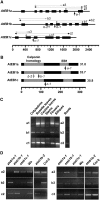
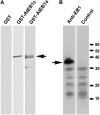
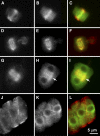
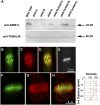
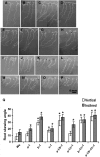
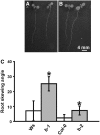
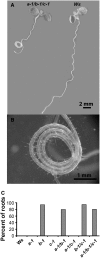
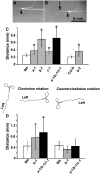
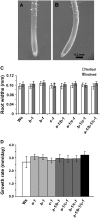
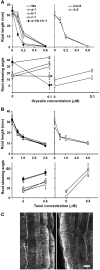
Microtubules function in concert with associated proteins that modify microtubule behavior and/or transmit signals that effect changes in growth. To better understand how microtubules and their associated proteins influence growth, we analyzed one family of microtubule-associated proteins, the END BINDING1 (EB1) proteins, in Arabidopsis thaliana (EB1a, EB1b, and EB1c). We find that antibodies directed against EB1 proteins colocalize with microtubules in roots, an observation that confirms previous reports using EB1-GFP fusions. We also find that T-DNA insertion mutants with reduced expression from EB1 genes have roots that deviate toward the left on vertical or inclined plates. Mutant roots also exhibit extended horizontal growth before they bend downward after tracking around an obstacle or after a 90 degrees clockwise reorientation of the root. These observations suggest that leftward deviations in root growth may be the result of delayed responses to touch and/or gravity signals. Root lengths and widths are normal, indicating that the delay in bend formation is not due to changes in the overall rate of growth. In addition, the genotype with the most severe defects responds to low doses of microtubule inhibitors in a manner indistinguishable from the wild type, indicating that microtubule integrity is not a major contributor to the leftward deviations in mutant root growth.
Figures
Similar articles
-
The microtubule plus-end tracking proteins SPR1 and EB1b interact to maintain polar cell elongation and directional organ growth in Arabidopsis.Plant Cell. 2014 Nov;26(11):4409-25. doi: 10.1105/tpc.114.131482. Epub 2014 Nov 18. Plant Cell. 2014. PMID: 25415978 Free PMC article.
-
Nuclear-localized subtype of end-binding 1 protein regulates spindle organization in Arabidopsis.J Cell Sci. 2010 Feb 1;123(Pt 3):451-9. doi: 10.1242/jcs.062703. Epub 2010 Jan 12. J Cell Sci. 2010. PMID: 20067996
-
The microtubule associated protein END BINDING 1 represses root responses to mechanical cues.Plant Sci. 2012 May;187:1-9. doi: 10.1016/j.plantsci.2012.01.010. Epub 2012 Jan 30. Plant Sci. 2012. PMID: 22404827
-
The Arabidopsis sku6/spiral1 gene encodes a plus end-localized microtubule-interacting protein involved in directional cell expansion.Plant Cell. 2004 Jun;16(6):1506-20. doi: 10.1105/tpc.020644. Epub 2004 May 21. Plant Cell. 2004. PMID: 15155883 Free PMC article.
-
Microtubule organization and microtubule-associated proteins in plant cells.Int Rev Cell Mol Biol. 2014;312:1-52. doi: 10.1016/B978-0-12-800178-3.00001-4. Int Rev Cell Mol Biol. 2014. PMID: 25262237 Review.
Cited by
-
A Cycloheximide-Sensitive Step in Transverse Microtubule Array Patterning.Plant Physiol. 2018 Oct;178(2):684-698. doi: 10.1104/pp.18.00672. Epub 2018 Aug 28. Plant Physiol. 2018. PMID: 30154175 Free PMC article.
-
EB1 and EB3 regulate microtubule minus end organization and Golgi morphology.J Cell Biol. 2017 Oct 2;216(10):3179-3198. doi: 10.1083/jcb.201701024. Epub 2017 Aug 16. J Cell Biol. 2017. PMID: 28814570 Free PMC article.
-
Two populations of cytoplasmic dynein contribute to spindle positioning in C. elegans embryos.J Cell Biol. 2017 Sep 4;216(9):2777-2793. doi: 10.1083/jcb.201607038. Epub 2017 Jul 24. J Cell Biol. 2017. PMID: 28739679 Free PMC article.
-
The role of 26S proteasome-dependent proteolysis in the formation and restructuring of microtubule networks.Plant Signal Behav. 2012 Oct 1;7(10):1289-95. doi: 10.4161/psb.21543. Epub 2012 Aug 20. Plant Signal Behav. 2012. PMID: 22902696 Free PMC article. Review.
-
Inhibition of phospholipase C disrupts cytoskeletal organization and gravitropic growth in Arabidopsis roots.Planta. 2010 Oct;232(5):1263-79. doi: 10.1007/s00425-010-1256-0. Epub 2010 Aug 29. Planta. 2010. PMID: 20803215
References
-
- Abe, T., and Hashimoto, T. (2005). Altered microtubule dynamics by expression of modified α-tubulin protein causes right-handed helical growth in transgenic Arabidopsis plants. Plant J. 43 191–204. - PubMed
-
- Alonso, J.M., et al. (2003). Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301 653–657. - PubMed
-
- Bienz, M. (2001). Spindles cotton on to junctions, APC and EB1. Nat. Cell Biol. 3 E1–E3. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

