EMBRYONIC FLOWER1 participates in polycomb group-mediated AG gene silencing in Arabidopsis
- PMID: 18281509
- PMCID: PMC2276442
- DOI: 10.1105/tpc.106.049957
EMBRYONIC FLOWER1 participates in polycomb group-mediated AG gene silencing in Arabidopsis
Abstract
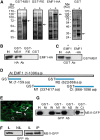
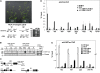
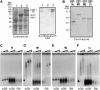
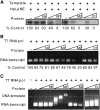
Polycomb group (PcG)-mediated gene silencing is a common developmental strategy used to maintain stably inherited repression of target genes and involves different protein complexes known as Polycomb-repressive complexes (PRCs). In animals, the two best-characterized PcG complexes are PRC1 and PRC2. In this report, we demonstrate that the plant-specific protein EMBRYONIC FLOWER1 (EMF1) functions in maintaining the repression of the flower homeotic gene AGAMOUS (AG) during vegetative development in Arabidopsis thaliana by acting in concert with the EMF2 complex, a putative equivalent of Drosophila melanogaster PRC2. We show that AG regulatory sequences are required for its ectopic expression in both emf1 and emf2 mutants and that EMF2 is required for trimethylation of histone 3 lysine 27 on the AG chromatin. We found that EMF1 interacts directly with AG and that this interaction depends on the presence of EMF2. Together with the finding of EMF1 interference with transcription in vitro, these results suggest that EMF1 enables transcriptional repression of AG after the action of the putative EMF2 complex. Our data indicate that EMF1 plays a PRC1-like role in the PcG-mediated floral repression mechanism.
Figures


Similar articles
-
Epigenetic regulation of gene programs by EMF1 and EMF2 in Arabidopsis.Plant Physiol. 2010 Feb;152(2):516-28. doi: 10.1104/pp.109.143495. Epub 2009 Sep 25. Plant Physiol. 2010. PMID: 19783648 Free PMC article.
-
Kicking against the PRCs - A Domesticated Transposase Antagonises Silencing Mediated by Polycomb Group Proteins and Is an Accessory Component of Polycomb Repressive Complex 2.PLoS Genet. 2015 Dec 7;11(12):e1005660. doi: 10.1371/journal.pgen.1005660. eCollection 2015 Dec. PLoS Genet. 2015. PMID: 26642436 Free PMC article.
-
EMF1 and PRC2 cooperate to repress key regulators of Arabidopsis development.PLoS Genet. 2012;8(3):e1002512. doi: 10.1371/journal.pgen.1002512. Epub 2012 Mar 22. PLoS Genet. 2012. PMID: 22457632 Free PMC article.
-
Polycomb-mediated gene silencing in Arabidopsis thaliana.Mol Cells. 2014 Dec 31;37(12):841-50. doi: 10.14348/molcells.2014.0249. Epub 2014 Nov 20. Mol Cells. 2014. PMID: 25410906 Free PMC article. Review.
-
Current understanding of plant Polycomb group proteins and the repressive histone H3 Lysine 27 trimethylation.Biochem Soc Trans. 2020 Aug 28;48(4):1697-1706. doi: 10.1042/BST20200192. Biochem Soc Trans. 2020. PMID: 32725200 Review.
Cited by
-
A core subunit of Polycomb repressive complex 1 is broadly conserved in function but not primary sequence.Proc Natl Acad Sci U S A. 2012 May 1;109(18):E1063-71. doi: 10.1073/pnas.1118678109. Epub 2012 Apr 18. Proc Natl Acad Sci U S A. 2012. PMID: 22517748 Free PMC article.
-
Regulation by polycomb and trithorax group proteins in Arabidopsis.Arabidopsis Book. 2010;8:e0128. doi: 10.1199/tab.0128. Epub 2010 May 8. Arabidopsis Book. 2010. PMID: 22303254 Free PMC article.
-
The Harbinger transposon-derived gene PANDA epigenetically coordinates panicle number and grain size in rice.Plant Biotechnol J. 2022 Jun;20(6):1154-1166. doi: 10.1111/pbi.13799. Epub 2022 Mar 16. Plant Biotechnol J. 2022. PMID: 35239255 Free PMC article.
-
LHP1 Could Act as an Activator and a Repressor of Transcription in Plants.Front Plant Sci. 2017 Nov 28;8:2041. doi: 10.3389/fpls.2017.02041. eCollection 2017. Front Plant Sci. 2017. PMID: 29234344 Free PMC article. Review.
-
The domesticated transposase ALP2 mediates formation of a novel Polycomb protein complex by direct interaction with MSI1, a core subunit of Polycomb Repressive Complex 2 (PRC2).PLoS Genet. 2020 May 28;16(5):e1008681. doi: 10.1371/journal.pgen.1008681. eCollection 2020 May. PLoS Genet. 2020. PMID: 32463832 Free PMC article.
References
-
- Alvarez-Venegas, R., Pien, S., Sadder, M., Witmer, X., Grossniklaus, U., and Avramova, Z. (2003). ATX-1, an Arabidopsis homolog of trithorax, activates flower homeotic genes. Curr. Biol. 13 627–637. - PubMed
-
- Bastow, R., Mylne, J.S., Lister, C., Lippman, Z., Martienssen, R.A., and Dean, C. (2004). Vernalization requires epigenetic silencing of FLC by histone methylation. Nature 427 164–167. - PubMed
-
- Bowler, C., Benvenuto, G., Laflamme, P., Molino, D., Probst, A.V., Tariq, M., and Paszkowski, J. (2004). Chromatin techniques for plant cells. Plant J. 39 776–789. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

