Fuzzy association rules for biological data analysis: a case study on yeast
- PMID: 18284669
- PMCID: PMC2277399
- DOI: 10.1186/1471-2105-9-107
Fuzzy association rules for biological data analysis: a case study on yeast
Abstract
Background: Last years' mapping of diverse genomes has generated huge amounts of biological data which are currently dispersed through many databases. Integration of the information available in the various databases is required to unveil possible associations relating already known data. Biological data are often imprecise and noisy. Fuzzy set theory is specially suitable to model imprecise data while association rules are very appropriate to integrate heterogeneous data.
Results: In this work we propose a novel fuzzy methodology based on a fuzzy association rule mining method for biological knowledge extraction. We apply this methodology over a yeast genome dataset containing heterogeneous information regarding structural and functional genome features. A number of association rules have been found, many of them agreeing with previous research in the area. In addition, a comparison between crisp and fuzzy results proves the fuzzy associations to be more reliable than crisp ones.
Conclusion: An integrative approach as the one carried out in this work can unveil significant knowledge which is currently hidden and dispersed through the existing biological databases. It is shown that fuzzy association rules can model this knowledge in an intuitive way by using linguistic labels and few easy-understandable parameters.
Figures
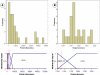
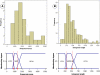
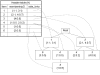


Similar articles
-
Biomedical application of fuzzy association rules for identifying breast cancer biomarkers.Med Biol Eng Comput. 2012 Sep;50(9):981-90. doi: 10.1007/s11517-012-0914-8. Epub 2012 May 24. Med Biol Eng Comput. 2012. PMID: 22622817
-
Protein superfamily classification using fuzzy rule-based classifier.IEEE Trans Nanobioscience. 2009 Mar;8(1):92-9. doi: 10.1109/TNB.2009.2016484. Epub 2009 Mar 21. IEEE Trans Nanobioscience. 2009. PMID: 19307166
-
Incremental fuzzy mining of gene expression data for gene function prediction.IEEE Trans Biomed Eng. 2011 May;58(5):1246-52. doi: 10.1109/TBME.2010.2047724. Epub 2010 Apr 15. IEEE Trans Biomed Eng. 2011. PMID: 20403777
-
Uncertainty of data, fuzzy membership functions, and multilayer perceptrons.IEEE Trans Neural Netw. 2005 Jan;16(1):10-23. doi: 10.1109/TNN.2004.836200. IEEE Trans Neural Netw. 2005. PMID: 15732386
-
Fuzzy OLAP association rules mining-based modular reinforcement learning approach for multiagent systems.IEEE Trans Syst Man Cybern B Cybern. 2005 Apr;35(2):326-38. doi: 10.1109/tsmcb.2004.843278. IEEE Trans Syst Man Cybern B Cybern. 2005. PMID: 15828660
Cited by
-
An intuitionistic approach to scoring DNA sequences against transcription factor binding site motifs.BMC Bioinformatics. 2010 Nov 8;11:551. doi: 10.1186/1471-2105-11-551. BMC Bioinformatics. 2010. PMID: 21059262 Free PMC article.
-
Mining Association Rules among Gene Functions in Clusters of Similar Gene Expression Maps.IEEE Int Conf Bioinform Biomed Workshops. 2009 Nov 1;2009:254-259. doi: 10.1109/BIBMW.2009.5332104. IEEE Int Conf Bioinform Biomed Workshops. 2009. PMID: 25635265 Free PMC article.
-
Biomedical application of fuzzy association rules for identifying breast cancer biomarkers.Med Biol Eng Comput. 2012 Sep;50(9):981-90. doi: 10.1007/s11517-012-0914-8. Epub 2012 May 24. Med Biol Eng Comput. 2012. PMID: 22622817
-
CisMiner: genome-wide in-silico cis-regulatory module prediction by fuzzy itemset mining.PLoS One. 2014 Sep 30;9(9):e108065. doi: 10.1371/journal.pone.0108065. eCollection 2014. PLoS One. 2014. PMID: 25268582 Free PMC article.
-
Data- and expert-driven rule induction and filtering framework for functional interpretation and description of gene sets.J Biomed Semantics. 2017 Jun 26;8(1):23. doi: 10.1186/s13326-017-0129-x. J Biomed Semantics. 2017. PMID: 28651634 Free PMC article.
References
-
- Kanehisa M, Bork P. Bioinformatics in the post-sequence era. Nature Genet. 2003;33:305–310. - PubMed
-
- Narayanan A, Keedwell EC, Olsson B. Artificial intelligence techniques for bioinformatics. Appl Bioinf. 2002;1:191–222. - PubMed
-
- Bhaskar H, Hoyle D, Singh S. Machine learning in bioinformatics: A brief survey and recommendations for practitioners. Computers in Biology and Medicine. 2005;36:1104–1125. Epub 2005 Oct 13. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

