FOXP3 promoter demethylation reveals the committed Treg population in humans
- PMID: 18286169
- PMCID: PMC2238816
- DOI: 10.1371/journal.pone.0001612
FOXP3 promoter demethylation reveals the committed Treg population in humans
Abstract
Background: Naturally occurring thymus derived regulatory T cells (Tregs) are central in the maintenance of self-tolerance. The transcription factor FOXP3 is crucial for the suppressive activity of Tregs and is considered the most specific marker for this population. However, human non regulatory T cells upregulate FOXP3 transiently upon activation which calls for other means to identify the Treg population. Since epigenetic mechanisms are involved in the establishment of stable gene expression patterns during cell differentiation, we hypothesized that the methylation profile of the FOXP3 promoter would allow the distinction of truly committed Tregs.
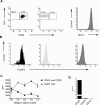
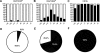
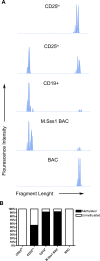
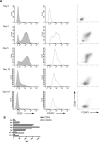
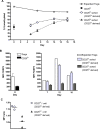
Methodology/principal findings: Human CD4(+)CD25(hi) Tregs displayed a demethylated FOXP3 promoter (1.4%+/-0.95% SEM methylated) in contrast to CD4(+)CD25(lo) T cells which were partially methylated (27.9%+/-7.1%). Furthermore, stimulated CD4(+)CD25(lo) T cells transiently expressed FOXP3 but remained partially methylated, suggesting promoter methylation as a mechanism for regulation of stable FOXP3 expression and Treg commitment. In addition, transient FOXP3 expressing cells exhibited suppressive abilities that correlate to the methylation status of the FOXP3 promoter. As an alternative to bisulphite sequencing, we present a restriction enzyme based screening method for the identification of committed Tregs and apply this method to evaluate the effect of various culturing conditions. We show that a partial demethylation occurs in long-term cultures after activation, whereas the addition of TGF-beta and/or IL-10 does not induce any additional change in methylation level.
Conclusions/significance: The unique FOXP3 promoter methylation profile in Tregs suggests that a demethylated pattern is a prerequisite for stable FOXP3 expression and suppressive phenotype. Presently, FOXP3 is used to identify Tregs in several human diseases and there are future implications for adoptive Treg transfer in immunotherapy. In these settings there is a need to distinguish true Tregs from transiently FOXP3(+) activated T cells. The screening method we present allows this distinction and enables the identification of cells suitable for in vitro expansions and clinical use.
Conflict of interest statement
Figures

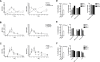
Similar articles
-
Isolation strategies of regulatory T cells for clinical trials: phenotype, function, stability, and expansion capacity.Exp Hematol. 2011 Dec;39(12):1152-60. doi: 10.1016/j.exphem.2011.08.010. Epub 2011 Aug 22. Exp Hematol. 2011. PMID: 21864487
-
T regulatory cells in cord blood--FOXP3 demethylation as reliable quantitative marker.PLoS One. 2010 Oct 12;5(10):e13267. doi: 10.1371/journal.pone.0013267. PLoS One. 2010. PMID: 20967272 Free PMC article.
-
Non-small-cell lung cancer-induced immunosuppression by increased human regulatory T cells via Foxp3 promoter demethylation.Cancer Immunol Immunother. 2016 May;65(5):587-99. doi: 10.1007/s00262-016-1825-6. Epub 2016 Mar 21. Cancer Immunol Immunother. 2016. PMID: 27000869 Free PMC article.
-
Epigenetic mechanisms of regulation of Foxp3 expression.Blood. 2009 Oct 29;114(18):3727-35. doi: 10.1182/blood-2009-05-219584. Epub 2009 Jul 29. Blood. 2009. PMID: 19641188 Free PMC article. Review.
-
Induction and maintenance of regulatory T cells by transcription factors and epigenetic modifications.J Autoimmun. 2017 Sep;83:113-121. doi: 10.1016/j.jaut.2017.07.002. Epub 2017 Jul 11. J Autoimmun. 2017. PMID: 28709726 Review.
Cited by
-
The role of regulatory T cells in atopic dermatitis.Curr Probl Dermatol. 2011;41:112-124. doi: 10.1159/000323305. Epub 2011 May 12. Curr Probl Dermatol. 2011. PMID: 21576952 Free PMC article. Review.
-
Activation of aryl hydrocarbon receptor (AhR) leads to reciprocal epigenetic regulation of FoxP3 and IL-17 expression and amelioration of experimental colitis.PLoS One. 2011;6(8):e23522. doi: 10.1371/journal.pone.0023522. Epub 2011 Aug 15. PLoS One. 2011. PMID: 21858153 Free PMC article.
-
1,25-Dihydroxyvitamin D3 and its analog TX527 promote a stable regulatory T cell phenotype in T cells from type 1 diabetes patients.PLoS One. 2014 Oct 3;9(10):e109194. doi: 10.1371/journal.pone.0109194. eCollection 2014. PLoS One. 2014. PMID: 25279717 Free PMC article.
-
Lkb1 maintains Treg cell lineage identity.Nat Commun. 2017 Jun 16;8:15876. doi: 10.1038/ncomms15876. Nat Commun. 2017. PMID: 28621313 Free PMC article.
-
Treg Cells and Epigenetic Regulation.Adv Exp Med Biol. 2021;1278:95-114. doi: 10.1007/978-981-15-6407-9_6. Adv Exp Med Biol. 2021. PMID: 33523445
References
-
- Fontenot JD, Gavin MA, Rudensky AY. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat Immunol. 2003;4:330–336. - PubMed
-
- Fontenot JD, Rasmussen JP, Williams LM, Dooley JL, Farr AG, et al. Regulatory T cell lineage specification by the forkhead transcription factor foxp3. Immunity. 2005;22:329–341. - PubMed
-
- Hori S, Nomura T, Sakaguchi S. Control of regulatory T cell development by the transcription factor Foxp3. Science. 2003;299:1057–1061. - PubMed
-
- Yagi H, Nomura T, Nakamura K, Yamazaki S, Kitawaki T, et al. Crucial role of FOXP3 in the development and function of human CD25+CD4+ regulatory T cells. Int Immunol. 2004;16:1643–1656. - PubMed
-
- Gambineri E, Torgerson TR, Ochs HD. Immune dysregulation, polyendocrinopathy, enteropathy, and X-linked inheritance (IPEX), a syndrome of systemic autoimmunity caused by mutations of FOXP3, a critical regulator of T-cell homeostasis. Curr Opin Rheumatol. 2003;15:430–435. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

