Regulation of chondrocytic gene expression by biomechanical signals
- PMID: 18304028
- PMCID: PMC4967411
- DOI: 10.1615/critreveukargeneexpr.v18.i2.30
Regulation of chondrocytic gene expression by biomechanical signals
Abstract
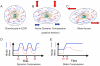
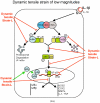
Cartilage is a mechanosensitive tissue, which means that it can perceive and respond to biomechanical signals. Despite the known importance of biomechanical signals in the etiopathogenesis of arthritic diseases and their effectiveness in joint restoration, little is understood about their actions at the cellular level. Recent molecular approaches have revealed that specific biomechanical stimuli and cell interactions generate intracellular signals that are powerful inducers or suppressors of proinflammatory and reparative genes in chondrocytes. Biomechanical signals are perceived by cartilage in magnitude-, frequency-, and time-dependent manners. Static and dynamic biomechanical forces of high magnitudes induce proinflammatory genes and inhibit matrix synthesis. Contrarily, dynamic biomechanical signals of low/physiologic magnitudes are potent antiinflammatory signals that inhibit interleukin-1beta (IL-1beta)-induced proinflammatory gene transcription and abrogate IL-1beta/tumor necrosis factor-alpha-induced inhibition of matrix synthesis. Recent studies have identified nuclear factor-kB (NF-kB) transcription factors as key regulators of biomechanical signal-mediated proinflammatory and antiinflammatory actions. These signals intercept multiple steps in the NF-kappaB signaling cascade to regulate cytokine gene expression. Taken together, these findings provide insight into how biomechanical signals regulate inflammatory and reparative gene transcription, underscoring their potential in enhancing the ability of chondrocytes to curb inflammation in diseased joints.
Figures


References
-
- Grodzinsky AJ, Levenston ME, Jin M, Frank EH. Cartilage tissue remodeling in response to mechanical forces. Annu Rev Biomed Eng. 2000;2:691–713. - PubMed
-
- Park S, Hung CT, Ateshian GA. Mechanical response of bovine articular cartilage under dynamic unconfined compression loading at physiological stress levels. Osteoarthritis Cartilage. 2004;12(1):65–73. - PubMed
-
- Fehrenbacher A, Steck E, Rickert M, Roth W, Richter W. Rapid regulation of collagen but not metalloproteinase 1, 3, 13, 14 and tissue inhibitor of metalloproteinase 1, 2, 3 expression in response to mechanical loading of cartilage explants in vitro. Arch Biochem Biophys. 2003;410(1):39–47. - PubMed
-
- Fitzgerald JB, Jin M, Grodzinsky AJ. Shear and compression differentially regulate clusters of functionally related temporal transcription patterns in cartilage tissue. J Biol Chem. 2006;281(34):24095–103. - PubMed
-
- Giannoni P, Siegrist M, Hunziker EB, Wong M. The mechanosensitivity of cartilage oligomeric matrix protein (COMP) Biorheology. 2003;40(1–3):101–9. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

