A complete collection of single-gene deletion mutants of Acinetobacter baylyi ADP1
- PMID: 18319726
- PMCID: PMC2290942
- DOI: 10.1038/msb.2008.10
A complete collection of single-gene deletion mutants of Acinetobacter baylyi ADP1
Abstract
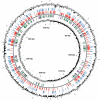
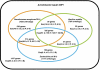
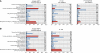
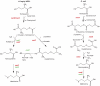
We have constructed a collection of single-gene deletion mutants for all dispensable genes of the soil bacterium Acinetobacter baylyi ADP1. A total of 2594 deletion mutants were obtained, whereas 499 (16%) were not, and are therefore candidate essential genes for life on minimal medium. This essentiality data set is 88% consistent with the Escherichia coli data set inferred from the Keio mutant collection profiled for growth on minimal medium, while 80% of the orthologous genes described as essential in Pseudomonas aeruginosa are also essential in ADP1. Several strategies were undertaken to investigate ADP1 metabolism by (1) searching for discrepancies between our essentiality data and current metabolic knowledge, (2) comparing this essentiality data set to those from other organisms, (3) systematic phenotyping of the mutant collection on a variety of carbon sources (quinate, 2-3 butanediol, glucose, etc.). This collection provides a new resource for the study of gene function by forward and reverse genetic approaches and constitutes a robust experimental data source for systems biology approaches.
Figures




References
-
- Abd El-Haleem D (2003) Acinetobacter: Environmental and Biotechnological Applications. Af J Biotechnol 2: 71–74
-
- Alaminos M, Ramos JL (2001) The methionine biosynthetic pathway from homoserine in Pseudomonas putida involves the metW, metX, metZ, metH and metE gene products. Arch Microbiol 176: 151–154 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

