Cue1p is an activator of Ubc7p E2 activity in vitro and in vivo
- PMID: 18321851
- PMCID: PMC2442330
- DOI: 10.1074/jbc.M801122200
Cue1p is an activator of Ubc7p E2 activity in vitro and in vivo
Abstract
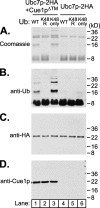
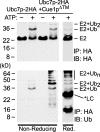
Ubc7p is a ubiquitin-conjugating enzyme (E2) that functions with endoplasmic reticulum (ER)-resident ubiquitin ligases (E3s) to promote endoplasmic reticulum-associated degradation (ERAD). Ubc7p only functions in ERAD if bound to the ER surface by Cue1p, a membrane-anchored ER protein. The role of Cue1p was thought to involve passive concentration of Ubc7p at the surface of the ER. However, our biochemical studies of Ubc7p suggested that Cue1p may, in addition, stimulate Ubc7p E2 activity. We have tested this idea and found it to be true both in vitro and in vivo. Ubc7p bound to the soluble domain of Cue1p showed strongly enhanced in vitro ubiquitination activity, both in the presence and absence of E3. Cue1p also enhanced Ubc7p function in vivo, and this activation was separable from the established ER-anchoring role of Cue1p. Finally, we tested in vivo activation of Ubc7p by Cue1p in an assay independent of the ER membrane and ERAD. A chimeric E2 linking Ubc7p to the Cdc34p/Ubc3p localization domain complemented the cdc34-2 TS phenotype, and co-expression of the soluble Cue1p domain enhanced complementation by this chimeric Ubc7p E2. These studies reveal a previously unobserved stimulation of Ubc7p E2 activity by Cue1p that is critical for full ERAD and that functions independently of the well known Cue1p anchoring function. Moreover, it suggests a previously unappreciated mode for regulation of E2s by Cue1p-like interacting partners.
Figures



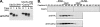
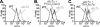
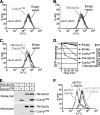


Similar articles
-
A Ubc7p-binding domain in Cue1p activates ER-associated protein degradation.J Cell Sci. 2009 May 1;122(Pt 9):1374-81. doi: 10.1242/jcs.044255. Epub 2009 Apr 14. J Cell Sci. 2009. PMID: 19366730 Free PMC article.
-
A structurally unique E2-binding domain activates ubiquitination by the ERAD E2, Ubc7p, through multiple mechanisms.Mol Cell. 2013 May 23;50(4):516-27. doi: 10.1016/j.molcel.2013.04.004. Epub 2013 May 9. Mol Cell. 2013. PMID: 23665230 Free PMC article.
-
Ubiquitin binding by a CUE domain regulates ubiquitin chain formation by ERAD E3 ligases.Mol Cell. 2013 May 23;50(4):528-39. doi: 10.1016/j.molcel.2013.04.005. Epub 2013 May 9. Mol Cell. 2013. PMID: 23665229
-
Endoplasmic reticulum degradation: reverse protein flow of no return.FASEB J. 1997 Dec;11(14):1227-33. doi: 10.1096/fasebj.11.14.9409541. FASEB J. 1997. PMID: 9409541 Review.
-
Ubiquitin ligases, critical mediators of endoplasmic reticulum-associated degradation.Semin Cell Dev Biol. 2007 Dec;18(6):770-9. doi: 10.1016/j.semcdb.2007.09.002. Epub 2007 Sep 8. Semin Cell Dev Biol. 2007. PMID: 17950636 Free PMC article. Review.
Cited by
-
The exportomer: the peroxisomal receptor export machinery.Cell Mol Life Sci. 2013 Apr;70(8):1393-411. doi: 10.1007/s00018-012-1136-9. Epub 2012 Sep 15. Cell Mol Life Sci. 2013. PMID: 22983384 Free PMC article. Review.
-
Protein quality control as a strategy for cellular regulation: lessons from ubiquitin-mediated regulation of the sterol pathway.Chem Rev. 2009 Apr;109(4):1561-74. doi: 10.1021/cr800544v. Chem Rev. 2009. PMID: 19243134 Free PMC article. Review. No abstract available.
-
The Unfolded Protein Response, Degradation from Endoplasmic Reticulum and Cancer.Genes Cancer. 2010 Jul 1;1(7):764-778. doi: 10.1177/1947601910383011. Genes Cancer. 2010. PMID: 21331300 Free PMC article.
-
HECT and RING finger families of E3 ubiquitin ligases at a glance.J Cell Sci. 2012 Feb 1;125(Pt 3):531-7. doi: 10.1242/jcs.091777. J Cell Sci. 2012. PMID: 22389392 Free PMC article. Review. No abstract available.
-
Enzymatic Logic of Ubiquitin Chain Assembly.Front Physiol. 2019 Jul 5;10:835. doi: 10.3389/fphys.2019.00835. eCollection 2019. Front Physiol. 2019. PMID: 31333493 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

