The molecular evolution of four anti-malarial immune genes in the Anopheles gambiae species complex
- PMID: 18325105
- PMCID: PMC2288592
- DOI: 10.1186/1471-2148-8-79
The molecular evolution of four anti-malarial immune genes in the Anopheles gambiae species complex
Abstract
Background: If the insect innate immune system is to be used as a potential blocking step in transmission of malaria, then it will require targeting one or a few genes with highest relevance and ease of manipulation. The problem is to identify and manipulate those of most importance to malaria infection without the risk of decreasing the mosquito's ability to stave off infections by microbes in general. Molecular evolution methodologies and concepts can help identify such genes. Within the setting of a comparative molecular population genetic and phylogenetic framework, involving six species of the Anopheles gambiae complex, we investigated whether a set of four pre-selected immunity genes (gambicin, NOS, Rel2 and FBN9) might have evolved under selection pressure imposed by the malaria parasite.
Results: We document varying levels of polymorphism within and divergence between the species, in all four genes. Introgression and the sharing of ancestral polymorphisms, two processes that have been documented in the past, were verified in this study in all four studied genes. These processes appear to affect each gene in different ways and to different degrees. However, there is no evidence of positive selection acting on these genes.
Conclusion: Considering the results presented here in concert with previous studies, genes that interact directly with the Plasmodium parasite, and play little or no role in defense against other microbes, are probably the most likely candidates for a specific adaptive response against P. falciparum. Furthermore, since it is hard to establish direct evidence linking the adaptation of any candidate gene to P. falciparum infection, a comparative framework allowing at least an indirect link should be provided. Such a framework could be achieved, if a similar approach like the one involved here, was applied to all other anopheline complexes that transmit P. falciparum malaria.
Figures
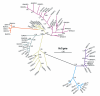
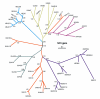
Similar articles
-
Polymorphisms in Anopheles gambiae immune genes associated with natural resistance to Plasmodium falciparum.PLoS Pathog. 2010 Sep 16;6(9):e1001112. doi: 10.1371/journal.ppat.1001112. PLoS Pathog. 2010. PMID: 20862317 Free PMC article.
-
The Anopheles FBN9 immune factor mediates Plasmodium species-specific defense through transgenic fat body expression.Dev Comp Immunol. 2017 Feb;67:257-265. doi: 10.1016/j.dci.2016.09.012. Epub 2016 Sep 22. Dev Comp Immunol. 2017. PMID: 27667688
-
Caspar controls resistance to Plasmodium falciparum in diverse anopheline species.PLoS Pathog. 2009 Mar;5(3):e1000335. doi: 10.1371/journal.ppat.1000335. Epub 2009 Mar 13. PLoS Pathog. 2009. PMID: 19282971 Free PMC article.
-
Comparative and functional genomics of the innate immune system in the malaria vector Anopheles gambiae.Immunol Rev. 2004 Apr;198:127-48. doi: 10.1111/j.0105-2896.2004.0127.x. Immunol Rev. 2004. PMID: 15199960 Review.
-
Innate immunity in the malaria vector Anopheles gambiae: comparative and functional genomics.J Exp Biol. 2004 Jul;207(Pt 15):2551-63. doi: 10.1242/jeb.01066. J Exp Biol. 2004. PMID: 15201288 Review.
Cited by
-
Anopheles immune genes and amino acid sites evolving under the effect of positive selection.PLoS One. 2010 Jan 26;5(1):e8885. doi: 10.1371/journal.pone.0008885. PLoS One. 2010. PMID: 20126662 Free PMC article.
-
Rodent malaria-resistant strains of the mosquito, Anopheles gambiae, have slower population growth than -susceptible strains.BMC Evol Biol. 2009 Apr 20;9:76. doi: 10.1186/1471-2148-9-76. BMC Evol Biol. 2009. PMID: 19379508 Free PMC article.
-
Single-nucleotide polymorphisms for high-throughput genotyping of Anopheles arabiensis in East and southern Africa.J Med Entomol. 2012 Mar;49(2):307-15. doi: 10.1603/me11113. J Med Entomol. 2012. PMID: 22493848 Free PMC article.
-
No evidence for positive selection at two potential targets for malaria transmission-blocking vaccines in Anopheles gambiae s.s.Infect Genet Evol. 2013 Jun;16:87-92. doi: 10.1016/j.meegid.2013.01.006. Epub 2013 Jan 26. Infect Genet Evol. 2013. PMID: 23357581 Free PMC article.
-
High amino acid diversity and positive selection at a putative coral immunity gene (tachylectin-2).BMC Evol Biol. 2010 May 19;10:150. doi: 10.1186/1471-2148-10-150. BMC Evol Biol. 2010. PMID: 20482872 Free PMC article.
References
-
- Takken W, Boëte C. Ecological aspects for application of genetically modified mosquitoes. Vol. 2. Dordrecht, Netherlands: Kluwer Academic Publishers; 2003. An introduction to ecological challenges concerning the use of genetically-modified mosquitoes for disease control; pp. 9–12.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

