Quantifying genes and transcripts to assess the in situ physiology of "Dehalococcoides" spp. in a trichloroethene-contaminated groundwater site
- PMID: 18326677
- PMCID: PMC2394903
- DOI: 10.1128/AEM.02199-07
Quantifying genes and transcripts to assess the in situ physiology of "Dehalococcoides" spp. in a trichloroethene-contaminated groundwater site
Abstract
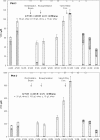
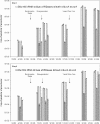
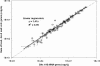
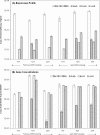
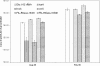
Quantitative PCR (qPCR) was coupled with reverse transcription (RT) to analyze both gene copy numbers and transcripts of the 16S rRNA gene and three reductive dehalogenase (RDase) genes (tceA, vcrA, and bvcA) as biomarkers of "Dehalococcoides" spp. in the groundwater of a trichloroethene-dense nonaqueous-phase liquid site at Fort Lewis, WA, that was sequentially subjected to biostimulation and bioaugmentation. Dehalococcoides cells carrying the tceA, vcrA, and bvcA genes were indigenous to the site. The sum of the three identified RDase gene copy numbers closely correlated to 16S rRNA gene copy numbers throughout the biostimulation and bioaugmentation activity, suggesting that these RDase genes represented the major Dehalococcoides metabolic functions at this site. Biomarker quantification revealed an overall increase of more than 3 orders of magnitude in the total Dehalococcoides population through the 1-year monitoring period (spanning biostimulation and bioaugmentation), and measurement of the respective RDase gene concentrations indicated different growth dynamics among Dehalococcoides cells. The Dehalococcoides cells containing the tceA gene consistently lagged behind other Dehalococcoides cells in population numbers and made up less than 5% of the total Dehalococcoides population, whereas the vcrA- and bvcA-containing cells represented the dominant fractions. Quantification of transcripts in groundwater samples verified that the 16S rRNA gene and the bvcA and vcrA genes were consistently highly expressed in all samples examined, while the tceA transcripts were detected inconsistently, suggesting a less active physiological state of the cells with this gene. The production of vinyl chloride and ethene toward the end of treatment supported the physiological activity of the bvcA- and vcrA-carrying cells. A clone library of the expressed RDase genes in field samples produced with degenerate primers revealed the expression of two putative RDase genes that were not previously monitored with RT-qPCR. The level of abundance of one of the putative RDase genes (FtL-RDase-1638) identified in the cDNA clone library tracked closely in field samples with abundance of the bvcA gene, suggesting that the FtL-RDase-1638 gene was likely colocated in genomes containing the bvcA gene. Overall, results from this study demonstrate that quantification of biomarker dynamics at field sites can provide useful information about the in situ physiology of Dehalococcoides strains and their associated activity.
Figures
Similar articles
-
Correlation of Dehalococcoides 16S rRNA and chloroethene-reductive dehalogenase genes with geochemical conditions in chloroethene-contaminated groundwater.Appl Environ Microbiol. 2010 Feb;76(3):843-50. doi: 10.1128/AEM.01482-09. Epub 2009 Dec 11. Appl Environ Microbiol. 2010. PMID: 20008170 Free PMC article.
-
GeneCARD-FISH: detection of tceA and vcrA reductive dehalogenase genes in Dehalococcoides mccartyi by fluorescence in situ hybridization.J Microbiol Methods. 2015 Mar;110:27-32. doi: 10.1016/j.mimet.2015.01.005. Epub 2015 Jan 14. J Microbiol Methods. 2015. PMID: 25595619
-
Oxygen effect on Dehalococcoides viability and biomarker quantification.Environ Sci Technol. 2008 Aug 1;42(15):5718-26. doi: 10.1021/es703227g. Environ Sci Technol. 2008. PMID: 18754499
-
Real-time PCR quantification of Dehalococcoides populations: methods and applications.J Microbiol Methods. 2008 Jan;72(1):1-11. doi: 10.1016/j.mimet.2007.11.005. Epub 2007 Nov 19. J Microbiol Methods. 2008. PMID: 18077025 Review.
-
The little bacteria that can - diversity, genomics and ecophysiology of 'Dehalococcoides' spp. in contaminated environments.Microb Biotechnol. 2010 Jul;3(4):389-402. doi: 10.1111/j.1751-7915.2009.00147.x. Epub 2009 Sep 4. Microb Biotechnol. 2010. PMID: 21255338 Free PMC article. Review.
Cited by
-
Molecular techniques in the biotechnological fight against halogenated compounds in anoxic environments.Microb Biotechnol. 2012 May;5(3):347-67. doi: 10.1111/j.1751-7915.2011.00313.x. Epub 2011 Nov 9. Microb Biotechnol. 2012. PMID: 22070763 Free PMC article. Review.
-
Detection of Organohalide-Respiring Enzyme Biomarkers at a Bioaugmented TCE-Contaminated Field Site.Front Microbiol. 2019 Jun 27;10:1433. doi: 10.3389/fmicb.2019.01433. eCollection 2019. Front Microbiol. 2019. PMID: 31316484 Free PMC article.
-
Global transcriptomic and proteomic responses of Dehalococcoides ethenogenes strain 195 to fixed nitrogen limitation.Appl Environ Microbiol. 2012 Mar;78(5):1424-36. doi: 10.1128/AEM.06792-11. Epub 2011 Dec 16. Appl Environ Microbiol. 2012. PMID: 22179257 Free PMC article.
-
In situ TCE degradation mediated by complex dehalorespiring communities during biostimulation processes.Microb Biotechnol. 2012 Sep;5(5):642-53. doi: 10.1111/j.1751-7915.2012.00339.x. Epub 2012 Mar 21. Microb Biotechnol. 2012. PMID: 22432919 Free PMC article.
-
Functional Genes and Bacterial Communities During Organohalide Respiration of Chloroethenes in Microcosms of Multi-Contaminated Groundwater.Front Microbiol. 2019 Feb 12;10:89. doi: 10.3389/fmicb.2019.00089. eCollection 2019. Front Microbiol. 2019. PMID: 30809199 Free PMC article.
References
-
- Adrian, L., U. Szewzyk, J. Wecke, and H. Görisch. 2000. Bacterial dehalorespiration with chlorinated benzenes. Nature 408:580-583. - PubMed
-
- Becker, J. G. 2006. A modeling study and implications of competition between Dehalococcoides ethenogenes and other tetrachloroethene-respiring bacteria. Environ. Sci. Technol. 40:4473-4480. - PubMed
-
- Daprato, R. C., F. E. Löffler, and J. B. Hughes. 2007. Comparative analysis of three tetrachloroethene to ethene halorespiring consortia suggests functional redundancy. Environ. Sci. Technol. 41:2261-2269. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

