Labeling of unique sequences in double-stranded DNA at sites of vicinal nicks generated by nicking endonucleases
- PMID: 18344522
- PMCID: PMC2373456
- DOI: 10.1093/nar/gkn107
Labeling of unique sequences in double-stranded DNA at sites of vicinal nicks generated by nicking endonucleases
Abstract
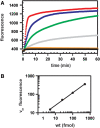
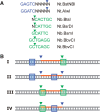
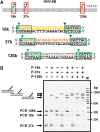
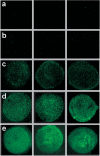
We describe a new approach for labeling of unique sequences within dsDNA under nondenaturing conditions. The method is based on the site-specific formation of vicinal nicks, which are created by nicking endonucleases (NEases) at specified DNA sites on the same strand within dsDNA. The oligomeric segment flanked by both nicks is then substituted, in a strand displacement reaction, by an oligonucleotide probe that becomes covalently attached to the target site upon subsequent ligation. Monitoring probe hybridization and ligation reactions by electrophoretic mobility retardation assay, we show that selected target sites can be quantitatively labeled with excellent sequence specificity. In these experiments, predominantly probes carrying a target-independent 3' terminal sequence were employed. At target labeling, thus a branched DNA structure known as 3'-flap DNA is obtained. The single-stranded terminus in 3'-flap DNA is then utilized to prime the replication of an externally supplied ssDNA circle in a rolling circle amplification (RCA) reaction. In model experiments with samples comprised of genomic lambda-DNA and human herpes virus 6 type B (HHV-6B) DNA, we have used our labeling method in combination with surface RCA as reporter system to achieve both high sequence specificity of dsDNA targeting and high sensitivity of detection. The method can find applications in sensitive and specific detection of viral duplex DNA.
Figures



Similar articles
-
Separate sites for binding and nicking of bacteriophage lambda DNA by terminase.Proc Natl Acad Sci U S A. 1983 Feb;80(4):955-9. doi: 10.1073/pnas.80.4.955. Proc Natl Acad Sci U S A. 1983. PMID: 6302676 Free PMC article.
-
Amplification of target-specific, ligation-dependent circular probe.Gene. 1998 May 12;211(2):277-85. doi: 10.1016/s0378-1119(98)00113-9. Gene. 1998. PMID: 9602151
-
Ultrasensitive assay based on a combined cascade amplification by nicking-mediated rolling circle amplification and symmetric strand-displacement amplification.Anal Chim Acta. 2019 Jan 24;1047:172-178. doi: 10.1016/j.aca.2018.10.004. Epub 2018 Oct 19. Anal Chim Acta. 2019. PMID: 30567647
-
Natural and engineered nicking endonucleases--from cleavage mechanism to engineering of strand-specificity.Nucleic Acids Res. 2011 Jan;39(1):1-18. doi: 10.1093/nar/gkq742. Epub 2010 Aug 30. Nucleic Acids Res. 2011. PMID: 20805246 Free PMC article. Review.
-
Amplification of circularizable probes for the detection of target nucleic acids and proteins.Clin Chim Acta. 2006 Jan;363(1-2):61-70. doi: 10.1016/j.cccn.2005.05.039. Epub 2005 Aug 24. Clin Chim Acta. 2006. PMID: 16122721 Review.
Cited by
-
Targeting individual subunits of the FokI restriction endonuclease to specific DNA strands.Nucleic Acids Res. 2009 Apr;37(7):2105-15. doi: 10.1093/nar/gkp046. Epub 2009 Feb 17. Nucleic Acids Res. 2009. PMID: 19223323 Free PMC article.
-
Oriented Soft DNA Curtains for Single-Molecule Imaging.Langmuir. 2021 Mar 23;37(11):3428-3437. doi: 10.1021/acs.langmuir.1c00066. Epub 2021 Mar 9. Langmuir. 2021. PMID: 33689355 Free PMC article.
-
Electrochemiluminescent determination of the activity of uracil-DNA glycosylase: Combining nicking enzyme assisted signal amplification and catalyzed hairpin assembly.Mikrochim Acta. 2019 Feb 15;186(3):179. doi: 10.1007/s00604-019-3280-5. Mikrochim Acta. 2019. PMID: 30771006
-
Efficient preparation of internally modified single-molecule constructs using nicking enzymes.Nucleic Acids Res. 2011 Feb;39(3):e15. doi: 10.1093/nar/gkq1004. Epub 2010 Nov 10. Nucleic Acids Res. 2011. PMID: 21071409 Free PMC article.
-
Simultaneous single-molecule measurements of phage T7 replisome composition and function reveal the mechanism of polymerase exchange.Proc Natl Acad Sci U S A. 2011 Mar 1;108(9):3584-9. doi: 10.1073/pnas.1018824108. Epub 2011 Jan 18. Proc Natl Acad Sci U S A. 2011. PMID: 21245349 Free PMC article.
References
-
- Ghosh I, Stains CI, Ooi AT, Segal DJ. Direct detection of double-stranded DNA: molecular methods and applications for DNA diagnostics. Mol. Biosyst. 2006;2:551–560. - PubMed
-
- Zelphati O, Liang X, Hobart P, Felgner PL. Gene chemistry: functionally and conformationally intact fluorescent plasmid DNA. Hum. Gene Ther. 1999;10:15–24. - PubMed
-
- Fechter EJ, Olenyuk B, Dervan PB. Sequence-specific fluorescence detection of DNA by polyamide-thiazole orange conjugates. J. Am. Chem. Soc. 2005;127:16685–16691. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

