piggyBac can bypass DNA synthesis during cut and paste transposition
- PMID: 18354502
- PMCID: PMC2323262
- DOI: 10.1038/emboj.2008.41
piggyBac can bypass DNA synthesis during cut and paste transposition
Abstract
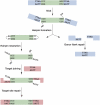
DNA synthesis is considered a defining feature in the movement of transposable elements. In determining the mechanism of piggyBac transposition, an insect transposon that is being increasingly used for genome manipulation in a variety of systems including mammalian cells, we have found that DNA synthesis can be avoided during piggyBac transposition, both at the donor site following transposon excision and at the insertion site following transposon integration. We demonstrate that piggyBac transposon excision occurs through the formation of transient hairpins on the transposon ends and that piggyBac target joining occurs by the direct attack of the 3'OH transposon ends on to the target DNA. This is the same strategy for target joining used by the members of DDE superfamily of transposases and retroviral integrases. Analysis of mutant piggyBac transposases in vitro and in vivo using a piggyBac transposition system we have established in Saccharomyces cerevisiae suggests that piggyBac transposase is a member of the DDE superfamily of recombinases, an unanticipated result because of the lack of sequence similarity between piggyBac and DDE family of recombinases.
Figures
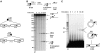



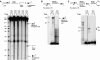
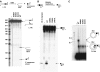
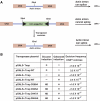
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

