Receptor tyrosine phosphatases regulate birth order-dependent axonal fasciculation and midline repulsion during development of the Drosophila mushroom body
- PMID: 18356078
- PMCID: PMC2435377
- DOI: 10.1016/j.mcn.2008.01.015
Receptor tyrosine phosphatases regulate birth order-dependent axonal fasciculation and midline repulsion during development of the Drosophila mushroom body
Abstract
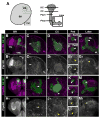
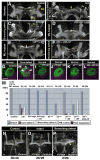
Receptor tyrosine phosphatases (RPTPs) are required for axon guidance during embryonic development in Drosophila. Here we examine the roles of four RPTPs during development of the larval mushroom body (MB). MB neurons extend axons into parallel tracts known as the peduncle and lobes. The temporal order of neuronal birth is reflected in the organization of axons within these tracts. Axons of the youngest neurons, known as core fibers, extend within a single bundle at the center, while those of older neurons fill the outer layers. RPTPs are selectively expressed on the core fibers of the MB. Ptp10D and Ptp69D regulate segregation of the young axons into a single core bundle. Ptp69D signaling is required for axonal extension beyond the peduncle. Lar and Ptp69D are necessary for the axonal branching decisions that create the lobes. Avoidance of the brain midline by extending medial lobe axons involves signaling through Lar.
Figures




Similar articles
-
Redundancy and compensation in axon guidance: genetic analysis of the Drosophila Ptp10D/Ptp4E receptor tyrosine phosphatase subfamily.Neural Dev. 2008 Jan 31;3:3. doi: 10.1186/1749-8104-3-3. Neural Dev. 2008. PMID: 18237413 Free PMC article.
-
The unfulfilled gene is required for the development of mushroom body neuropil in Drosophila.Neural Dev. 2010 Feb 1;5:4. doi: 10.1186/1749-8104-5-4. Neural Dev. 2010. PMID: 20122139 Free PMC article.
-
The receptor protein tyrosine phosphatase PTP69D antagonizes Abl tyrosine kinase to guide axons in Drosophila.Mech Dev. 2008 Mar-Apr;125(3-4):247-56. doi: 10.1016/j.mod.2007.11.005. Epub 2007 Nov 22. Mech Dev. 2008. PMID: 18160268 Free PMC article.
-
Protein tyrosine phosphatases and neural development.Bioessays. 1998 Jun;20(6):463-72. doi: 10.1002/(SICI)1521-1878(199806)20:6<463::AID-BIES4>3.0.CO;2-N. Bioessays. 1998. PMID: 9699458 Review.
-
Receptor tyrosine phosphatases in axon growth and guidance.Curr Opin Neurobiol. 2001 Feb;11(1):95-102. doi: 10.1016/s0959-4388(00)00179-3. Curr Opin Neurobiol. 2001. PMID: 11179878 Review.
Cited by
-
Structure-function analyses of tyrosine phosphatase PTP69D in giant fiber synapse formation of Drosophila.Mol Cell Neurosci. 2015 Jan;64:24-31. doi: 10.1016/j.mcn.2014.11.002. Epub 2014 Nov 26. Mol Cell Neurosci. 2015. PMID: 25433167 Free PMC article.
-
Protein tyrosine phosphatase receptor type O (Ptpro) regulates cerebellar formation during zebrafish development through modulating Fgf signaling.Cell Mol Life Sci. 2013 Jul;70(13):2367-81. doi: 10.1007/s00018-013-1259-7. Epub 2013 Jan 30. Cell Mol Life Sci. 2013. PMID: 23361036 Free PMC article.
-
Developmental changes in expression, subcellular distribution, and function of Drosophila N-cadherin, guided by a cell-intrinsic program during neuronal differentiation.Dev Biol. 2012 Jun 15;366(2):204-17. doi: 10.1016/j.ydbio.2012.04.006. Epub 2012 Apr 19. Dev Biol. 2012. PMID: 22542600 Free PMC article.
-
The making of the Drosophila mushroom body.Front Physiol. 2023 Jan 13;14:1091248. doi: 10.3389/fphys.2023.1091248. eCollection 2023. Front Physiol. 2023. PMID: 36711013 Free PMC article. Review.
-
Glial Derived TGF-β Instructs Axon Midline Stopping.Front Mol Neurosci. 2019 Sep 27;12:232. doi: 10.3389/fnmol.2019.00232. eCollection 2019. Front Mol Neurosci. 2019. PMID: 31611773 Free PMC article.
References
-
- Babu K, Bahri S, Alphey L, Chia W. Bifocal and PP1 interaction regulates targeting of the R-cell growth cone in Drosophila. Developmental biology. 2005;288:372–386. - PubMed
-
- Clandinin TR, Lee CH, Herman T, Lee RC, Yang AY, Ovasapyan S, Zipursky SL. Drosophila LAR regulates R1–R6 and R7 target specificity in the visual system. Neuron. 2001;32:237–248. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

