Dissection of the essential steps for condensin accumulation at kinetochores and rDNAs during fission yeast mitosis
- PMID: 18362178
- PMCID: PMC2290841
- DOI: 10.1083/jcb.200708170
Dissection of the essential steps for condensin accumulation at kinetochores and rDNAs during fission yeast mitosis
Abstract
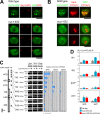
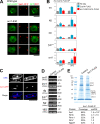
The condensin complex has a fundamental role in chromosome dynamics. In this study, we report that accumulation of Schizosaccharomyces pombe condensin at mitotic kinetochores and ribosomal DNAs (rDNAs) occurs in multiple steps and is necessary for normal segregation of the sister kinetochores and rDNAs. Nuclear entry of condensin at the onset of mitosis requires Cut15/importin alpha and Cdc2 phosphorylation. Ark1/aurora and Cut17/Bir1/survivin are needed to dock the condensin at both the kinetochores and rDNAs. Furthermore, proteins that are necessary to form the chromatin architecture of the kinetochores (Mis6, Cnp1, and Mis13) and rDNAs (Nuc1 and Acr1) are required for condensin to accumulate specifically at these sites. Acr1 (accumulation of condensin at rDNA 1) is an rDNA upstream sequence binding protein that physically interacts with Rrn5, Rrn11, Rrn7, and Spp27 and is required for the proper accumulation of Nuc1 at rDNAs. The mechanism of condensin accumulation at the kinetochores may be conserved, as human condensin II fails to accumulate at kinetochores in hMis6 RNA interference-treated cells.
Figures






Similar articles
-
Condensin-mediated chromosome organization in fission yeast.Curr Genet. 2016 Nov;62(4):739-743. doi: 10.1007/s00294-016-0601-7. Epub 2016 Apr 9. Curr Genet. 2016. PMID: 27061734 Free PMC article. Review.
-
Chromosome segregation: monopolin attracts condensin.Curr Biol. 2011 Aug 23;21(16):R634-6. doi: 10.1016/j.cub.2011.06.059. Curr Biol. 2011. PMID: 21855006 Free PMC article.
-
Condensin association with histone H2A shapes mitotic chromosomes.Nature. 2011 Jun 1;474(7352):477-83. doi: 10.1038/nature10179. Nature. 2011. PMID: 21633354
-
Condensin phosphorylated by the Aurora-B-like kinase Ark1 is continuously required until telophase in a mode distinct from Top2.J Cell Sci. 2011 Jun 1;124(Pt 11):1795-807. doi: 10.1242/jcs.078733. Epub 2011 May 3. J Cell Sci. 2011. PMID: 21540296
-
The regulation of chromosome segregation via centromere loops.Crit Rev Biochem Mol Biol. 2019 Aug;54(4):352-370. doi: 10.1080/10409238.2019.1670130. Epub 2019 Oct 1. Crit Rev Biochem Mol Biol. 2019. PMID: 31573359 Free PMC article. Review.
Cited by
-
Condensin-mediated chromosome organization in fission yeast.Curr Genet. 2016 Nov;62(4):739-743. doi: 10.1007/s00294-016-0601-7. Epub 2016 Apr 9. Curr Genet. 2016. PMID: 27061734 Free PMC article. Review.
-
Aurora B prevents chromosome arm separation defects by promoting telomere dispersion and disjunction.J Cell Biol. 2015 Mar 16;208(6):713-27. doi: 10.1083/jcb.201407016. J Cell Biol. 2015. PMID: 25778919 Free PMC article.
-
Inositol Pyrophosphate Kinase Asp1 Modulates Chromosome Segregation Fidelity and Spindle Function in Schizosaccharomyces pombe.Mol Cell Biol. 2016 Nov 28;36(24):3128-3140. doi: 10.1128/MCB.00330-16. Print 2016 Dec 15. Mol Cell Biol. 2016. PMID: 27697865 Free PMC article.
-
Condensin: Architect of mitotic chromosomes.Chromosome Res. 2009;17(2):131-44. doi: 10.1007/s10577-008-9009-7. Chromosome Res. 2009. PMID: 19308696 Review.
-
The ABCs of CENPs.Chromosoma. 2011 Oct;120(5):425-46. doi: 10.1007/s00412-011-0330-0. Epub 2011 Jul 13. Chromosoma. 2011. PMID: 21751032 Review.
References
-
- Aoki, K., Y. Nakaseko, K. Kinoshita, G. Goshima, and M. Yanagida. 2006. CDC2 phosphorylation of the fission yeast dis1 ensures accurate chromosome segregation. Curr. Biol. 16:1627–1635. - PubMed
-
- Aono, N., T. Sutani, T. Tomonaga, S. Mochida, and M. Yanagida. 2002. Cnd2 has dual roles in mitotic condensation and interphase. Nature. 417:197–202. - PubMed
-
- Appelgren, H., B. Kniola, and K. Ekwall. 2003. Distinct centromere domain structures with separate functions demonstrated in live fission yeast cells. J. Cell Sci. 116:4035–4042. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

