Distinct methylation changes at the IGF2-H19 locus in congenital growth disorders and cancer
- PMID: 18365005
- PMCID: PMC2268001
- DOI: 10.1371/journal.pone.0001849
Distinct methylation changes at the IGF2-H19 locus in congenital growth disorders and cancer
Abstract
Background: Differentially methylated regions (DMRs) are associated with many imprinted genes. In mice methylation at a DMR upstream of the H19 gene known as the Imprint Control region (IC1) is acquired in the male germline and influences the methylation status of DMRs 100 kb away in the adjacent Insulin-like growth factor 2 (Igf2) gene through long-range interactions. In humans, germline-derived or post-zygotically acquired imprinting defects at IC1 are associated with aberrant activation or repression of IGF2, resulting in the congenital growth disorders Beckwith-Wiedemann (BWS) and Silver-Russell (SRS) syndromes, respectively. In Wilms tumour and colorectal cancer, biallelic expression of IGF2 has been observed in association with loss of methylation at a DMR in IGF2. This DMR, known as DMR0, has been shown to be methylated on the silent maternal IGF2 allele presumably with a role in repression. The effect of IGF2 DMR0 methylation changes in the aetiology of BWS or SRS is unknown.
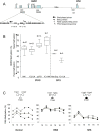
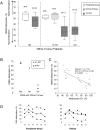
Methodology/principal findings: We analysed the methylation status of the DMR0 in BWS, SRS and Wilms tumour patients by conventional bisulphite sequencing and pyrosequencing. We show here that, contrary to previous reports, the IGF2 DMR0 is actually methylated on the active paternal allele in peripheral blood and kidney. This is similar to the IC1 methylation status and is inconsistent with the proposed silencing function of the maternal IGF2 allele. Beckwith-Wiedemann and Silver-Russell patients with IC1 methylation defects have similar methylation defects at the IGF2 DMR0, consistent with IC1 regulating methylation at IGF2 in cis. In Wilms tumour, however, methylation profiles of IC1 and IGF2 DMR0 are indicative of methylation changes occurring on both parental alleles rather than in cis.
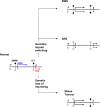
Conclusions/significance: These results support a model in which DMR0 and IC1 have opposite susceptibilities to global hyper and hypomethylation during tumorigenesis independent of the parent of origin imprint. In contrast, during embryogenesis DMR0 is methylated or demethylated according to the germline methylation imprint at the IC1, indicating different mechanisms of imprinting loss in neoplastic and non-neoplastic cells.
Conflict of interest statement
Figures
References
-
- Cooper WN, Luharia A, Evans GA, Raza H, Haire AC, et al. Molecular subtypes and phenotypic expression of Beckwith-Wiedemann syndrome. Eur J Hum Genet. 2005;13:1025–1032. - PubMed
-
- Delaval K, Wagschal A, Feil R. Epigenetic deregulation of imprinting in congenital diseases of aberrant growth. Bioessays. 2006;28:453–459. - PubMed
-
- Feinberg AP, Tycko B. The history of cancer epigenetics. Nat Rev Cancer. 2004;4:143–153. - PubMed
-
- Netchine I, Rossignol S, Dufourg MN, Azzi S, Rousseau A, et al. 11p15 imprinting center region 1 loss of methylation is a common and specific cause of typical Russell-Silver syndrome: clinical scoring system and epigenetic-phenotypic correlations. J Clin Endocrinol Metab. 2007;92:3148–3154. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

