Phosphorylation-independent regulation of the diguanylate cyclase WspR
- PMID: 18366254
- PMCID: PMC2270323
- DOI: 10.1371/journal.pbio.0060067
Phosphorylation-independent regulation of the diguanylate cyclase WspR
Abstract
Environmental signals that trigger bacterial pathogenesis and biofilm formation are mediated by changes in the level of cyclic dimeric guanosine monophosphate (c-di-GMP), a unique eubacterial second messenger. Tight regulation of cellular c-di-GMP concentration is governed by diguanylate cyclases and phosphodiesterases, which are responsible for its production and degradation, respectively. Here, we present the crystal structure of the diguanylate cyclase WspR, a conserved GGDEF domain-containing response regulator in Gram-negative bacteria, bound to c-di-GMP at an inhibitory site. Biochemical analyses revealed that feedback regulation involves the formation of at least three distinct oligomeric states. By switching from an active to a product-inhibited dimer via a tetrameric assembly, WspR utilizes a novel mechanism for modulation of its activity through oligomerization. Moreover, our data suggest that these enzymes can be activated by phosphodiesterases. Thus, in addition to the canonical pathways via phosphorylation of the regulatory domains, both product and enzyme concentration contribute to the coordination of c-di-GMP signaling. A structural comparison reveals resemblance of the oligomeric states to assemblies of GAF domains, widely used regulatory domains in signaling molecules conserved from archaea to mammals, suggesting a similar mechanism of regulation.
Conflict of interest statement
Figures

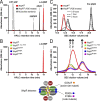
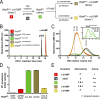

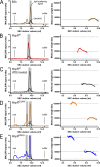
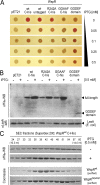


Comment in
-
Structural insight into a biofilm signaling molecule.PLoS Biol. 2008 Mar;6(3):e89. doi: 10.1371/journal.pbio.0060089. Epub 2008 Mar 25. PLoS Biol. 2008. PMID: 20076707 Free PMC article. No abstract available.
References
-
- Stock AM, Robinson VL, Goudreau PN. Two-component signal transduction. Annu Rev Biochem. 2000;69:183–215. - PubMed
-
- D'Argenio D, Miller S. Cyclic di-GMP as a bacterial second messenger. Microbiology. 2004;150:2497–2502. - PubMed
-
- Römling U, Gomelsky M, Galperin M. C-di-GMP: the dawning of a novel bacterial signalling system. Mol Microbiol. 2005;57:629–639. - PubMed
-
- Ross P, Mayer R, Weinhouse H, Amikam D, Huggirat Y, et al. The cyclic diguanylic acid regulatory system of cellulose synthesis in Acetobacter xylinum. Chemical synthesis and biological activity of cyclic nucleotide dimer, trimer, and phosphothioate derivatives. J Biol Chem. 1990;265:18933–18943. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

