Supervised learning method for the prediction of subcellular localization of proteins using amino acid and amino acid pair composition
- PMID: 18366605
- PMCID: PMC2386058
- DOI: 10.1186/1471-2164-9-S1-S16
Supervised learning method for the prediction of subcellular localization of proteins using amino acid and amino acid pair composition
Abstract
Background: Occurrence of protein in the cell is an important step in understanding its function. It is highly desirable to predict a protein's subcellular locations automatically from its sequence. Most studied methods for prediction of subcellular localization of proteins are signal peptides, the location by sequence homology, and the correlation between the total amino acid compositions of proteins. Taking amino-acid composition and amino acid pair composition into consideration helps improving the prediction accuracy.
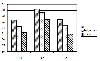
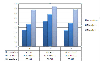
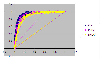
Results: We constructed a dataset of protein sequences from SWISS-PROT database and segmented them into 12 classes based on their subcellular locations. SVM modules were trained to predict the subcellular location based on amino acid composition and amino acid pair composition. Results were calculated after 10-fold cross validation. Radial Basis Function (RBF) outperformed polynomial and linear kernel functions. Total prediction accuracy reached to 71.8% for amino acid composition and 77.0% for amino acid pair composition. In order to observe the impact of number of subcellular locations we constructed two more datasets of nine and five subcellular locations. Total accuracy was further improved to 79.9% and 85.66%.
Conclusions: A new SVM based approach is presented based on amino acid and amino acid pair composition. Result shows that data simulation and taking more protein features into consideration improves the accuracy to a great extent. It was also noticed that the data set needs to be crafted to take account of the distribution of data in all the classes.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

