Interrogating domain-domain interactions with parsimony based approaches
- PMID: 18366803
- PMCID: PMC2358894
- DOI: 10.1186/1471-2105-9-171
Interrogating domain-domain interactions with parsimony based approaches
Abstract
Background: The identification and characterization of interacting domain pairs is an important step towards understanding protein interactions. In the last few years, several methods to predict domain interactions have been proposed. Understanding the power and the limitations of these methods is key to the development of improved approaches and better understanding of the nature of these interactions.
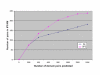
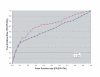
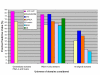
Results: Building on the previously published Parsimonious Explanation method (PE) to predict domain-domain interactions, we introduced a new Generalized Parsimonious Explanation (GPE) method, which (i) adjusts the granularity of the domain definition to the granularity of the input data set and (ii) permits domain interactions to have different costs. This allowed for preferential selection of the so-called "co-occurring domains" as possible mediators of interactions between proteins. The performance of both variants of the parsimony method are competitive to the performance of the top algorithms for this problem even though parsimony methods use less information than some of the other methods. We also examined possible enrichment of co-occurring domains and homo-domains among domain interactions mediating the interaction of proteins in the network. The corresponding study was performed by surveying domain interactions predicted by the GPE method as well as by using a combinatorial counting approach independent of any prediction method. Our findings indicate that, while there is a considerable propensity towards these special domain pairs among predicted domain interactions, this overrepresentation is significantly lower than in the iPfam dataset.
Conclusion: The Generalized Parsimonious Explanation approach provides a new means to predict and study domain-domain interactions. We showed that, under the assumption that all protein interactions in the network are mediated by domain interactions, there exists a significant deviation of the properties of domain interactions mediating interactions in the network from that of iPfam data.
Figures
Similar articles
-
An integrated approach to the prediction of domain-domain interactions.BMC Bioinformatics. 2006 May 25;7:269. doi: 10.1186/1471-2105-7-269. BMC Bioinformatics. 2006. PMID: 16725050 Free PMC article.
-
Discovering motif pairs at interaction sites from protein sequences on a proteome-wide scale.Bioinformatics. 2006 Apr 15;22(8):989-96. doi: 10.1093/bioinformatics/btl020. Epub 2006 Jan 29. Bioinformatics. 2006. PMID: 16446278
-
A domain-based approach to predict protein-protein interactions.BMC Bioinformatics. 2007 Jun 13;8:199. doi: 10.1186/1471-2105-8-199. BMC Bioinformatics. 2007. PMID: 17567909 Free PMC article.
-
Deciphering protein-protein interactions. Part II. Computational methods to predict protein and domain interaction partners.PLoS Comput Biol. 2007 Apr 27;3(4):e43. doi: 10.1371/journal.pcbi.0030043. PLoS Comput Biol. 2007. PMID: 17465672 Free PMC article. Review.
-
Deciphering protein-protein interactions. Part I. Experimental techniques and databases.PLoS Comput Biol. 2007 Mar 30;3(3):e42. doi: 10.1371/journal.pcbi.0030042. PLoS Comput Biol. 2007. PMID: 17397251 Free PMC article. Review. No abstract available.
Cited by
-
Protein-protein interaction networks (PPI) and complex diseases.Gastroenterol Hepatol Bed Bench. 2014 Winter;7(1):17-31. Gastroenterol Hepatol Bed Bench. 2014. PMID: 25436094 Free PMC article. Review.
-
IDDI: integrated domain-domain interaction and protein interaction analysis system.Proteome Sci. 2012 Jun 21;10 Suppl 1(Suppl 1):S9. doi: 10.1186/1477-5956-10-S1-S9. Proteome Sci. 2012. PMID: 22759586 Free PMC article.
-
PPIDomainMiner: Inferring domain-domain interactions from multiple sources of protein-protein interactions.PLoS Comput Biol. 2021 Aug 9;17(8):e1008844. doi: 10.1371/journal.pcbi.1008844. eCollection 2021 Aug. PLoS Comput Biol. 2021. PMID: 34370723 Free PMC article.
-
Inference of domain-disease associations from domain-protein, protein-disease and disease-disease relationships.BMC Syst Biol. 2016 Jan 11;10 Suppl 1(Suppl 1):4. doi: 10.1186/s12918-015-0247-y. BMC Syst Biol. 2016. PMID: 26818594 Free PMC article.
-
DOMINE: a comprehensive collection of known and predicted domain-domain interactions.Nucleic Acids Res. 2011 Jan;39(Database issue):D730-5. doi: 10.1093/nar/gkq1229. Epub 2010 Nov 27. Nucleic Acids Res. 2011. PMID: 21113022 Free PMC article.
References
-
- Uetz P, Giot L, Cagney G, Mansfield TA, Judson RS, Knight JR, Lockshon D, Narayan V, Srinivasan M, Pochart P, Qureshi-Emili A, Li Y, Godwin B, Conover D, Kalbfleisch T, Vijayadamodar G, Yang M, Johnston M, Fields S, Rothberg JM. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature. 2000;403:623–627. doi: 10.1038/35001009. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

