Proteomics analysis unravels the functional repertoire of coronavirus nonstructural protein 3
- PMID: 18367524
- PMCID: PMC2395186
- DOI: 10.1128/JVI.02631-07
Proteomics analysis unravels the functional repertoire of coronavirus nonstructural protein 3
Abstract
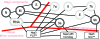
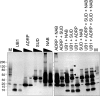
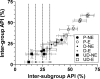
Severe acute respiratory syndrome (SARS) coronavirus infection and growth are dependent on initiating signaling and enzyme actions upon viral entry into the host cell. Proteins packaged during virus assembly may subsequently form the first line of attack and host manipulation upon infection. A complete characterization of virion components is therefore important to understanding the dynamics of early stages of infection. Mass spectrometry and kinase profiling techniques identified nearly 200 incorporated host and viral proteins. We used published interaction data to identify hubs of connectivity with potential significance for virion formation. Surprisingly, the hub with the most potential connections was not the viral M protein but the nonstructural protein 3 (nsp3), which is one of the novel virion components identified by mass spectrometry. Based on new experimental data and a bioinformatics analysis across the Coronaviridae, we propose a higher-resolution functional domain architecture for nsp3 that determines the interaction capacity of this protein. Using recombinant protein domains expressed in Escherichia coli, we identified two additional RNA-binding domains of nsp3. One of these domains is located within the previously described SARS-unique domain, and there is a nucleic acid chaperone-like domain located immediately downstream of the papain-like proteinase domain. We also identified a novel cysteine-coordinated metal ion-binding domain. Analyses of interdomain interactions and provisional functional annotation of the remaining, so-far-uncharacterized domains are presented. Overall, the ensemble of data surveyed here paint a more complete picture of nsp3 as a conserved component of the viral protein processing machinery, which is intimately associated with viral RNA in its role as a virion component.
Figures


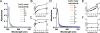


References
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215403-410. - PubMed
-
- Barabási, A. L., and Z. N. Oltvai. 2004. Network biology: understanding the cell's functional organization. Nat. Rev. Genet. 5101-113. - PubMed
-
- Bern, M., D. Goldberg, W. H. McDonald, and J. R. Yates III. 2004. Automatic quality assessment of peptide tandem mass spectra. Bioinformatics 20(Suppl. 1)i49-i54. - PubMed
-
- Bertini, I., and C. Luchinat. 1984. High spin cobalt(II) as a probe for the investigation of metalloproteins. Adv. Inorg. Biochem. 671-111. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

