Essential role of domain III of nonstructural protein 5A for hepatitis C virus infectious particle assembly
- PMID: 18369481
- PMCID: PMC2268006
- DOI: 10.1371/journal.ppat.1000035
Essential role of domain III of nonstructural protein 5A for hepatitis C virus infectious particle assembly
Abstract
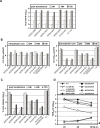
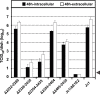
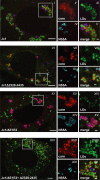
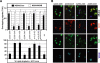
Persistent infection with the hepatitis C virus (HCV) is a major risk factor for the development of liver cirrhosis and hepatocellular carcinoma. With an estimated about 3% of the world population infected with this virus, the lack of a prophylactic vaccine and a selective therapy, chronic hepatitis C currently is a main indication for liver transplantation. The establishment of cell-based replication and virus production systems has led to first insights into the functions of HCV proteins. However, the role of nonstructural protein 5A (NS5A) in the viral replication cycle is so far not known. NS5A is a membrane-associated RNA-binding protein assumed to be involved in HCV RNA replication. Its numerous interactions with the host cell suggest that NS5A is also an important determinant for pathogenesis and persistence. In this study we show that NS5A is a key factor for the assembly of infectious HCV particles. We specifically identify the C-terminal domain III as the primary determinant in NS5A for particle formation. We show that both core and NS5A colocalize on the surface of lipid droplets, a proposed site for HCV particle assembly. Deletions in domain III of NS5A disrupting this colocalization abrogate infectious particle formation and lead to an enhanced accumulation of core protein on the surface of lipid droplets. Finally, we show that mutations in NS5A causing an assembly defect can be rescued by trans-complementation. These data provide novel insights into the production of infectious HCV and identify NS5A as a major determinant for HCV assembly. Since domain III of NS5A is one of the most variable regions in the HCV genome, the results suggest that viral isolates may differ in their level of virion production and thus in their level of fitness and pathogenesis.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Bialek SR, Terrault NA. The changing epidemiology and natural history of hepatitis C virus infection. Clin Liver Dis. 2006;10:697–715. - PubMed
-
- Gale M, Jr, Foy EM. Evasion of intracellular host defence by hepatitis C virus. Nature. 2005;436:939–945. - PubMed
-
- Bowen DG, Walker CM. Adaptive immune responses in acute and chronic hepatitis C virus infection. Nature. 2005;436:946–952. - PubMed
-
- De Francesco R, Migliaccio G. Challenges and successes in developing new therapies for hepatitis C. Nature. 2005;436:953–960. - PubMed
-
- Houghton M, Abrignani S. Prospects for a vaccine against the hepatitis C virus. Nature. 2005;436:961–966. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

