Mammalian long-chain acyl-CoA synthetases
- PMID: 18375835
- PMCID: PMC3377585
- DOI: 10.3181/0710-MR-287
Mammalian long-chain acyl-CoA synthetases
Abstract
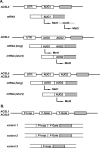
Acyl-CoA synthetase enzymes are essential for de novo lipid synthesis, fatty acid catabolism, and remodeling of membranes. Activation of fatty acids requires a two-step reaction catalyzed by these enzymes. In the first step, an acyl-AMP intermediate is formed from ATP. AMP is then exchanged with CoA to produce the activated acyl-CoA. The release of AMP in this reaction defines the superfamily of AMP-forming enzymes. The length of the carbon chain of the fatty acid species defines the substrate specificity for the different acyl-CoA synthetases (ACS). On this basis, five sub-families of ACS have been characterized. The purpose of this review is to report on the large family of mammalian long-chain acyl-CoA synthetases (ACSL), which activate fatty acids with chain lengths of 12 to 20 carbon atoms. Five genes and several isoforms generated by alternative splicing have been identified and limited information is available on their localization. The structure of these membrane proteins has not been solved for the mammalian ACSLs but homology to a bacterial form, whose structure has been determined, points at specific structural features that are important for these enzymes across species. The bacterial form acts as a dimer and has a conserved short motif, called the fatty acid Gate domain, that seems to determine substrate specificity. We will discuss the characterization and identification of the different spliced isoforms, draw attention to the inconsistencies and errors in their annotations, and their cellular localizations. These membrane proteins act on membrane-bound substrates probably as homo- and as heterodimer complexes but have often been expressed as single recombinant isoforms, apparently purified as monomers and tested in Triton X-100 micelles. We will argue that such studies have failed to provide an accurate assessment of the activity and of the distinct function of these enzymes in mammalian cells.
Figures







References
-
- Fujino T, Man-Jong K, Minekura H, Suzuki H, Yamamoto TT. Alternative translation initiation generates acyl-CoA synthetase 3 isoforms with heterogeneous amino termini. J Biochem (Tokyo) 1997;122:212–216. - PubMed
-
- Mashek DG, Bornfeldt KE, Coleman RA, Berger J, Bernlohr DA, Black P, DiRusso CC, Farber SA, Guo W, Hashimoto N, Khodiyar V, Kuypers FA, Maltais LJ, Nebert DW, Renieri A, Schaffer JE, Stahl A, Watkins PA, Vasiliou V, Yamamoto TT. Revised nomenclature for the mammalian long-chain acyl-CoA synthetase gene family. J Lipid Res. 2004;45:1958–1961. - PubMed
-
- Van Horn CG, Caviglia JM, Li LO, Wang S, Granger DA, Coleman RA. Characterization of recombinant long-chain rat acyl-CoA synthetase isoforms 3 and 6: identification of a novel variant of isoform 6. Biochemistry. 2005;44:1635–1642. - PubMed
-
- Hisanaga Y, Ago H, Nakagawa N, Hamada K, Ida K, Yamamoto M, Hori T, Arii Y, Sugahara M, Kuramitsu S, Yokoyama S, Miyano M. Structural basis of the substrate-specific two-step catalysis of long chain fatty acyl-CoA synthetase dimer. J Biol Chem. 2004;279:31717–31726. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

