A BBP-Mud2p heterodimer mediates branchpoint recognition and influences splicing substrate abundance in budding yeast
- PMID: 18375978
- PMCID: PMC2377449
- DOI: 10.1093/nar/gkn144
A BBP-Mud2p heterodimer mediates branchpoint recognition and influences splicing substrate abundance in budding yeast
Abstract
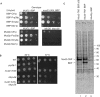
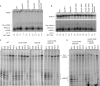
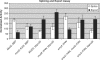
The 3' end of mammalian introns is marked by the branchpoint binding protein, SF1, and the U2AF65-U2AF35 heterodimer bound at an adjacent sequence. Baker's yeast has equivalent proteins, branchpoint binding protein (BBP) (SF1) and Mud2p (U2AF65), but lacks an obvious U2AF35 homolog, leaving open the question of whether another protein substitutes during spliceosome assembly. Gel filtration, affinity selection and mass spectrometry were used to show that rather than a U2AF65/U2AF35-like heterodimer, Mud2p forms a complex with BBP without a third (U2AF35-like) factor. Using mutants of MUD2 and BBP, we show that the BBP-Mud2p complex bridges partner-specific Prp39p, Mer1p, Clf1p and Smy2p two-hybrid interactions. In addition to inhibiting Mud2p association, the bbpDelta56 mutation impairs splicing, enhances pre-mRNA release from the nucleus, and similar to a mud2::KAN knockout, suppresses a lethal sub2::KAN mutation. Unexpectedly, rather than exacerbating bbpDelta56, the mud2::KAN mutation partially suppresses a pre-mRNA accumulation defect observed with bbpDelta56. We propose that a BBP-Mud2p heterodimer binds as a unit to the branchpoint in vivo and serves as a target for the Sub2p-DExD/H-box ATPase and for other splicing factors during spliceosome assembly. In addition, our results suggest the possibility that the Mud2p may enhance the turnover of pre-mRNA with impaired BBP-branchpoint association.
Figures



References
-
- Burge CB, Tuschl T, Sharp PA. Splicing of Precursors to mRNAs by the Spliceosome. In: Gesteland RF, Cech TR, Atkins JF, editors. The RNA World. 2nd. NY: Cold Spring Harbor Laboratory Press, Cold Spring Harbor; 1999.
-
- Abovich N, Rosbash M. Cross-intron bridging interactions in the yeast commitment complex are conserved in mammals. Cell. 1997;89:403–412. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

