The regulatory activity of microRNA* species has substantial influence on microRNA and 3' UTR evolution
- PMID: 18376413
- PMCID: PMC2698667
- DOI: 10.1038/nsmb.1409
The regulatory activity of microRNA* species has substantial influence on microRNA and 3' UTR evolution
Abstract
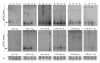
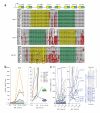
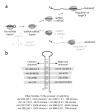
During microRNA (miRNA) biogenesis, one strand of a approximately 21-22-nucleotide RNA duplex is preferentially selected for entry into a silencing complex. The other strand, known as the miRNA* species, has typically been assumed to be a carrier strand. Here we show that, although Drosophila melanogaster miRNA* species are less abundant than their partners, they are often present at physiologically relevant levels and can associate with Argonaute proteins. Comparative genomic analyses revealed that >40% of miRNA* sequences resist nucleotide divergence across Drosophilid evolution, and at least half of these well-conserved miRNA* species select for conserved 3' untranslated region seed matches well above background noise. Finally, we validated the inhibitory activity of miRNA* species in both cultured cells and transgenic animals. These data broaden the reach of the miRNA regulatory network and suggest an important mechanism that diversifies miRNA function during evolution.
Figures




References
-
- Lai EC. MicroRNAs: runts of the genome assert themselves. Curr. Biol. 2003;13:R925–R936. - PubMed
-
- Bushati N, Cohen SM. MicroRNA functions. Annu. Rev. Cell Dev. Biol. 2007;23:175–205. - PubMed
-
- Lee Y, et al. The nuclear RNase III Drosha initiates microRNA processing. Nature. 2003;425:415–419. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

