The p12 domain is unstructured in a murine leukemia virus p12-CA(N) Gag construct
- PMID: 18382677
- PMCID: PMC2277328
- DOI: 10.1371/journal.pone.0001902
The p12 domain is unstructured in a murine leukemia virus p12-CA(N) Gag construct
Abstract
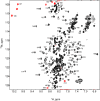
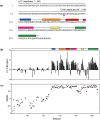
The Gag polyproteins of gammaretroviruses contain a conserved p12 domain between MA and CA that plays critical roles in virus assembly, reverse transcription and nuclear integration. Here we show using nuclear magnetic resonance, that p12 is unstructured in a Moloney murine leukemia virus (MMLV) Gag fragment that includes the N-terminal domain of CA (p12-CA(N)). Furthermore, no long range interactions were observed between the domains, as has been previously predicted. Flexibility appears to be a common feature of Gag "late" domains required for virus release during budding. Residues near the N-terminus of CA(N) that form a beta-hairpin in the mature CA protein are unfolded in p12-CA(N), consistent with proposals that hairpin formation helps trigger capsid assembly.
Conflict of interest statement
Figures
Similar articles
-
The N-terminus of murine leukaemia virus p12 protein is required for mature core stability.PLoS Pathog. 2014 Oct 30;10(10):e1004474. doi: 10.1371/journal.ppat.1004474. eCollection 2014 Oct. PLoS Pathog. 2014. PMID: 25356837 Free PMC article.
-
Repression of the Chromatin-Tethering Domain of Murine Leukemia Virus p12.J Virol. 2016 Nov 28;90(24):11197-11207. doi: 10.1128/JVI.01084-16. Print 2016 Dec 15. J Virol. 2016. PMID: 27707926 Free PMC article.
-
Critical Role of the Human T-Cell Leukemia Virus Type 1 Capsid N-Terminal Domain for Gag-Gag Interactions and Virus Particle Assembly.J Virol. 2018 Jun 29;92(14):e00333-18. doi: 10.1128/JVI.00333-18. Print 2018 Jul 15. J Virol. 2018. PMID: 29695435 Free PMC article.
-
Moloney Murine Leukemia Virus p12 Is Required for Histone Loading onto Retroviral DNAs.J Virol. 2021 Jul 12;95(15):e0049521. doi: 10.1128/JVI.00495-21. Epub 2021 Jul 12. J Virol. 2021. PMID: 34011543 Free PMC article.
-
Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly.EMBO J. 1998 Mar 16;17(6):1555-68. doi: 10.1093/emboj/17.6.1555. EMBO J. 1998. PMID: 9501077 Free PMC article.
Cited by
-
Hydrodynamic and Membrane Binding Properties of Purified Rous Sarcoma Virus Gag Protein.J Virol. 2015 Oct;89(20):10371-82. doi: 10.1128/JVI.01628-15. Epub 2015 Aug 5. J Virol. 2015. PMID: 26246573 Free PMC article.
-
Specific insertions of zinc finger domains into Gag-Pol yield engineered retroviral vectors with selective integration properties.Proc Natl Acad Sci U S A. 2010 Jul 13;107(28):12475-80. doi: 10.1073/pnas.1001402107. Epub 2010 Jun 28. Proc Natl Acad Sci U S A. 2010. PMID: 20616052 Free PMC article.
-
Murine leukemia virus p12 tethers the capsid-containing pre-integration complex to chromatin by binding directly to host nucleosomes in mitosis.PLoS Pathog. 2018 Jun 15;14(6):e1007117. doi: 10.1371/journal.ppat.1007117. eCollection 2018 Jun. PLoS Pathog. 2018. PMID: 29906285 Free PMC article.
-
Distinct Particle Morphologies Revealed through Comparative Parallel Analyses of Retrovirus-Like Particles.J Virol. 2016 Aug 26;90(18):8074-84. doi: 10.1128/JVI.00666-16. Print 2016 Sep 15. J Virol. 2016. PMID: 27356903 Free PMC article.
-
Nuclear Magnetic Resonance Spectroscopy to Study Virus Structure.Subcell Biochem. 2024;105:171-206. doi: 10.1007/978-3-031-65187-8_5. Subcell Biochem. 2024. PMID: 39738947 Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

