Cds1 controls the release of Cdc14-like phosphatase Flp1 from the nucleolus to drive full activation of the checkpoint response to replication stress in fission yeast
- PMID: 18385517
- PMCID: PMC2397296
- DOI: 10.1091/mbc.e07-08-0737
Cds1 controls the release of Cdc14-like phosphatase Flp1 from the nucleolus to drive full activation of the checkpoint response to replication stress in fission yeast
Abstract
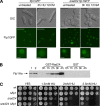
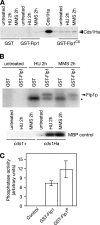
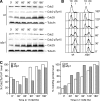
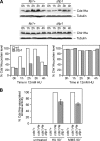
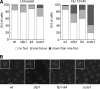
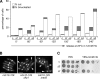
The Cdc14p-like phosphatase Flp1p (also known as Clp1p) is regulated by cell cycle-dependent changes in its subcellular localization. Flp1p is restricted to the nucleolus and spindle pole body until prophase, when it is dispersed throughout the nucleus, mitotic spindle, and medial ring. Once released, Flp1p antagonizes Cdc2p/cyclin activity by reverting Cdc2p-phosphorylation sites on Cdc25p. On replication stress, ataxia-telangiectasia mutated/ATM/Rad3-related kinase Rad3p activates Cds1p, which phosphorylates key proteins ensuring the stability of stalled DNA replication forks. Here, we show that replication stress induces changes in the subcellular localization of Flp1p in a checkpoint-dependent manner. Active Cds1p checkpoint kinase is required to release Flp1p into the nucleus. Consistently, a Flp1p mutant (flp1-9A) lacking all potential Cds1p phosphorylation sites fails to relocate in response to replication blocks and, similarly to cells lacking flp1 (Deltaflp1), presents defects in checkpoint response to replication stress. Deltaflp1 cells accumulate reduced levels of a less active Cds1p kinase in hydroxyurea (HU), indicating that nuclear Flp1p regulates Cds1p full activation. Consistently, Deltaflp1 and flp1-9A have an increased percentage of Rad22p-recombination foci during HU treatment. Together, our data show that by releasing Flp1p into the nucleus Cds1p checkpoint kinase modulates its own full activation during replication stress.
Figures



Similar articles
-
S. pombe FEAR protein orthologs are not required for release of Clp1/Flp1 phosphatase from the nucleolus during mitosis.J Cell Sci. 2006 Nov 1;119(Pt 21):4462-6. doi: 10.1242/jcs.03220. Epub 2006 Oct 10. J Cell Sci. 2006. PMID: 17032733
-
Replication checkpoint enforced by kinases Cds1 and Chk1.Science. 1998 May 8;280(5365):909-12. doi: 10.1126/science.280.5365.909. Science. 1998. PMID: 9572736
-
Multiple protein kinases influence the redistribution of fission yeast Clp1/Cdc14 phosphatase upon genotoxic stress.Mol Biol Cell. 2012 Oct;23(20):4118-28. doi: 10.1091/mbc.E12-06-0475. Epub 2012 Aug 23. Mol Biol Cell. 2012. PMID: 22918952 Free PMC article.
-
Flp1, a fission yeast orthologue of the s. cerevisiae CDC14 gene, is not required for cyclin degradation or rum1p stabilisation at the end of mitosis.J Cell Sci. 2001 Jul;114(Pt 14):2649-64. doi: 10.1242/jcs.114.14.2649. J Cell Sci. 2001. PMID: 11683392
-
Rad3 and Sty1 function in Schizosaccharomyces pombe: an integrated response to DNA damage and environmental stress?Mol Microbiol. 2008 Apr;68(2):246-54. doi: 10.1111/j.1365-2958.2008.06147.x. Mol Microbiol. 2008. PMID: 18366437 Review.
Cited by
-
Comprehensive proteomics analysis reveals new substrates and regulators of the fission yeast clp1/cdc14 phosphatase.Mol Cell Proteomics. 2013 May;12(5):1074-86. doi: 10.1074/mcp.M112.025924. Epub 2013 Jan 7. Mol Cell Proteomics. 2013. PMID: 23297348 Free PMC article.
-
Biochemical analyses reveal amino acid residues critical for cell cycle-dependent phosphorylation of human Cdc14A phosphatase by cyclin-dependent kinase 1.Sci Rep. 2018 Aug 8;8(1):11871. doi: 10.1038/s41598-018-30253-8. Sci Rep. 2018. PMID: 30089874 Free PMC article.
-
Orderly progression through S-phase requires dynamic ubiquitylation and deubiquitylation of PCNA.Sci Rep. 2016 May 6;6:25513. doi: 10.1038/srep25513. Sci Rep. 2016. PMID: 27151298 Free PMC article.
-
The Fission Yeast Cell Integrity Pathway: A Functional Hub for Cell Survival upon Stress and Beyond.J Fungi (Basel). 2021 Dec 30;8(1):32. doi: 10.3390/jof8010032. J Fungi (Basel). 2021. PMID: 35049972 Free PMC article. Review.
-
Cdc14 phosphatase contributes to cell wall integrity and pathogenesis in Candida albicans.Front Microbiol. 2023 Feb 16;14:1129155. doi: 10.3389/fmicb.2023.1129155. eCollection 2023. Front Microbiol. 2023. PMID: 36876065 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

