Synthetic lethal screens identify gene silencing processes in yeast and implicate the acetylated amino terminus of Sir3 in recognition of the nucleosome core
- PMID: 18391024
- PMCID: PMC2423298
- DOI: 10.1128/MCB.02050-07
Synthetic lethal screens identify gene silencing processes in yeast and implicate the acetylated amino terminus of Sir3 in recognition of the nucleosome core
Abstract
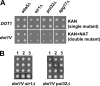
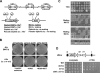
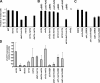
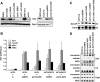
Dot1 methylates histone H3 lysine 79 (H3K79) on the nucleosome core and is involved in Sir protein-mediated silencing. Previous studies suggested that H3K79 methylation within euchromatin prevents nonspecific binding of the Sir proteins, which in turn facilitates binding of the Sir proteins in unmethylated silent chromatin. However, the mechanism by which the Sir protein binding is influenced by this modification is unclear. We performed genome-wide synthetic genetic array (SGA) analysis and identified interactions of DOT1 with SIR1 and POL32. The synthetic growth defects found by SGA analysis were attributed to the loss of mating type identity caused by a synthetic silencing defect. By using epistasis analysis, DOT1, SIR1, and POL32 could be placed in different pathways of silencing. Dot1 shared its silencing phenotypes with the NatA N-terminal acetyltransferase complex and the conserved N-terminal bromo adjacent homology (BAH) domain of Sir3 (a substrate of NatA). We classified all of these as affecting a common silencing process, and we show that mutations in this process lead to nonspecific binding of Sir3 to chromatin. Our results suggest that the BAH domain of Sir3 binds to histone H3K79 and that acetylation of the BAH domain is required for the binding specificity of Sir3 for nucleosomes unmethylated at H3K79.
Figures

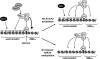
Similar articles
-
Structural basis of silencing: Sir3 BAH domain in complex with a nucleosome at 3.0 Å resolution.Science. 2011 Nov 18;334(6058):977-82. doi: 10.1126/science.1210915. Science. 2011. PMID: 22096199 Free PMC article.
-
Multiple histone modifications in euchromatin promote heterochromatin formation by redundant mechanisms in Saccharomyces cerevisiae.BMC Mol Biol. 2009 Jul 28;10:76. doi: 10.1186/1471-2199-10-76. BMC Mol Biol. 2009. PMID: 19638198 Free PMC article.
-
Structure and function of the Saccharomyces cerevisiae Sir3 BAH domain.Mol Cell Biol. 2006 Apr;26(8):3256-65. doi: 10.1128/MCB.26.8.3256-3265.2006. Mol Cell Biol. 2006. PMID: 16581798 Free PMC article.
-
Silent information regulator 3: the Goldilocks of the silencing complex.Genes Dev. 2010 Jan 15;24(2):115-22. doi: 10.1101/gad.1865510. Genes Dev. 2010. PMID: 20080949 Free PMC article. Review.
-
SIR-nucleosome interactions: structure-function relationships in yeast silent chromatin.Gene. 2013 Sep 15;527(1):10-25. doi: 10.1016/j.gene.2013.05.088. Epub 2013 Jun 18. Gene. 2013. PMID: 23791651 Review.
Cited by
-
Symmetry, asymmetry, and kinetics of silencing establishment in Saccharomyces cerevisiae revealed by single-cell optical assays.Proc Natl Acad Sci U S A. 2011 Jan 25;108(4):1209-16. doi: 10.1073/pnas.1018742108. Epub 2011 Jan 24. Proc Natl Acad Sci U S A. 2011. PMID: 21262833 Free PMC article.
-
Direct screening for chromatin status on DNA barcodes in yeast delineates the regulome of H3K79 methylation by Dot1.Elife. 2016 Dec 6;5:e18919. doi: 10.7554/eLife.18919. Elife. 2016. PMID: 27922451 Free PMC article.
-
Competition between Heterochromatic Loci Allows the Abundance of the Silencing Protein, Sir4, to Regulate de novo Assembly of Heterochromatin.PLoS Genet. 2015 Nov 20;11(11):e1005425. doi: 10.1371/journal.pgen.1005425. eCollection 2015 Nov. PLoS Genet. 2015. PMID: 26587833 Free PMC article.
-
The diverse functions of Dot1 and H3K79 methylation.Genes Dev. 2011 Jul 1;25(13):1345-58. doi: 10.1101/gad.2057811. Genes Dev. 2011. PMID: 21724828 Free PMC article. Review.
-
The 80th Threonine Residue of Histone H3 Is Important for Maintaining HM Silencing in Saccharomyces cerevisiae.J Microbiol Biotechnol. 2024 Jan 28;34(1):39-46. doi: 10.4014/jmb.2310.10031. Epub 2023 Nov 15. J Microbiol Biotechnol. 2024. PMID: 37957109 Free PMC article.
References
-
- Bi, X., Q. Yu, J. J. Sandmeier, and S. Elizondo. 2004. Regulation of transcriptional silencing in yeast by growth temperature. J. Mol. Biol. 344893-905. - PubMed
-
- Boone, C., H. Bussey, and B. J. Andrews. 2007. Exploring genetic interactions and networks with yeast. Nat. Rev. Genet. 8437-449. - PubMed
-
- Bose, M. E., K. H. McConnell, K. A. Gardner-Aukema, U. Muller, M. Weinreich, J. L. Keck, and C. A. Fox. 2004. The origin recognition complex and Sir4 protein recruit Sir1p to yeast silent chromatin through independent interactions requiring a common Sir1p domain. Mol. Cell. Biol. 24774-786. - PMC - PubMed
-
- Bostelman, L. J., A. M. Keller, A. M. Albrecht, A. Arat, and J. S. Thompson. 2007. Methylation of histone H3 lysine-79 by Dot1p plays multiple roles in the response to UV damage in Saccharomyces cerevisiae. DNA Repair (Amsterdam) 6383-395. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
