Identification of a potent synthetic FXR agonist with an unexpected mode of binding and activation
- PMID: 18391212
- PMCID: PMC2291122
- DOI: 10.1073/pnas.0710981105
Identification of a potent synthetic FXR agonist with an unexpected mode of binding and activation
Abstract
The farnesoid X receptor (FXR), a member of the nuclear hormone receptor family, plays important roles in the regulation of bile acid and cholesterol homeostasis, glucose metabolism, and insulin sensitivity. There is intense interest in understanding the mechanisms of FXR regulation and in developing pharmaceutically suitable synthetic FXR ligands that might be used to treat metabolic syndrome. We report here the identification of a potent FXR agonist (MFA-1) and the elucidation of the structure of this ligand in ternary complex with the human receptor and a coactivator peptide fragment using x-ray crystallography at 1.9-A resolution. The steroid ring system of MFA-1 binds with its D ring-facing helix 12 (AF-2) in a manner reminiscent of hormone binding to classical steroid hormone receptors and the reverse of the pose adopted by naturally occurring bile acids when bound to FXR. This binding mode appears to be driven by the presence of a carboxylate on MFA-1 that is situated to make a salt-bridge interaction with an arginine residue in the FXR-binding pocket that is normally used to neutralize bound bile acids. Receptor activation by MFA-1 differs from that by bile acids in that it relies on direct interactions between the ligand and residues in helices 11 and 12 and only indirectly involves a protonated histidine that is part of the activation trigger. The structure of the FXR:MFA-1 complex differs significantly from that of the complex with a structurally distinct agonist, fexaramine, highlighting the inherent plasticity of the receptor.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




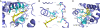

Similar articles
-
5Alpha-bile alcohols function as farnesoid X receptor antagonists.Biochem Biophys Res Commun. 2006 Jan 6;339(1):386-91. doi: 10.1016/j.bbrc.2005.11.027. Epub 2005 Nov 14. Biochem Biophys Res Commun. 2006. PMID: 16300737
-
Phosphorylation of farnesoid X receptor by protein kinase C promotes its transcriptional activity.Mol Endocrinol. 2008 Nov;22(11):2433-47. doi: 10.1210/me.2008-0092. Epub 2008 Aug 28. Mol Endocrinol. 2008. PMID: 18755856
-
Targeting farnesoid X receptor for liver and metabolic disorders.Trends Mol Med. 2007 Jul;13(7):298-309. doi: 10.1016/j.molmed.2007.06.001. Epub 2007 Jun 27. Trends Mol Med. 2007. PMID: 17588816 Review.
-
Conformationally constrained farnesoid X receptor (FXR) agonists: Naphthoic acid-based analogs of GW 4064.Bioorg Med Chem Lett. 2008 Aug 1;18(15):4339-43. doi: 10.1016/j.bmcl.2008.06.073. Epub 2008 Jun 28. Bioorg Med Chem Lett. 2008. PMID: 18621523
-
Bile salt excretory pump: biology and pathobiology.J Pediatr Gastroenterol Nutr. 2006 Jul;43 Suppl 1:S10-6. doi: 10.1097/01.mpg.0000226385.71859.5f. J Pediatr Gastroenterol Nutr. 2006. PMID: 16819395 Review.
Cited by
-
Identification of trisubstituted-pyrazol carboxamide analogs as novel and potent antagonists of farnesoid X receptor.Bioorg Med Chem. 2014 Jun 1;22(11):2919-38. doi: 10.1016/j.bmc.2014.04.014. Epub 2014 Apr 16. Bioorg Med Chem. 2014. PMID: 24775917 Free PMC article.
-
Molecular tuning of farnesoid X receptor partial agonism.Nat Commun. 2019 Jul 2;10(1):2915. doi: 10.1038/s41467-019-10853-2. Nat Commun. 2019. PMID: 31266946 Free PMC article.
-
Discovery of Farnesoid X Receptor Antagonists Based on a Library of Oleanolic Acid 3-O-Esters through Diverse Substituent Design and Molecular Docking Methods.Molecules. 2017 Apr 26;22(5):690. doi: 10.3390/molecules22050690. Molecules. 2017. PMID: 28445411 Free PMC article.
-
Exploring the binding diversity of intrinsically disordered proteins involved in one-to-many binding.Protein Sci. 2013 Mar;22(3):258-73. doi: 10.1002/pro.2207. Epub 2013 Jan 27. Protein Sci. 2013. PMID: 23233352 Free PMC article.
-
Discovery that theonellasterol a marine sponge sterol is a highly selective FXR antagonist that protects against liver injury in cholestasis.PLoS One. 2012;7(1):e30443. doi: 10.1371/journal.pone.0030443. Epub 2012 Jan 23. PLoS One. 2012. PMID: 22291955 Free PMC article.
References
-
- Goodwin B, et al. A regulatory cascade of the nuclear receptors FXR, SHP-1, and LRH-1 represses bile acid biosynthesis. Mol Cell. 2000;6:517–526. - PubMed
-
- Makishima M, et al. Identification of a nuclear receptor for bile acids. Science. 1999;284:1362–1365. - PubMed
-
- Parks DJ, et al. Bile acids: natural ligands for an orphan nuclear receptor. Science. 1999;284:1365–1368. - PubMed
-
- Wang H, et al. Endogenous bile acids are ligands for the nuclear receptor FXR/BAR. Mol Cell. 1999;3:543–553. - PubMed
-
- Lu TT, et al. Molecular basis for feedback regulation of bile acid synthesis by nuclear receptors. Mol Cell. 2000;6:507–515. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Chemical Information
Molecular Biology Databases

