Exploring structural variability in X-ray crystallographic models using protein local optimization by torsion-angle sampling
- PMID: 18391405
- PMCID: PMC2631124
- DOI: 10.1107/S090744490800070X
Exploring structural variability in X-ray crystallographic models using protein local optimization by torsion-angle sampling
Abstract
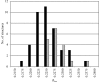
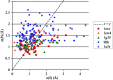
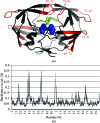
Modeling structural variability is critical for understanding protein function and for modeling reliable targets for in silico docking experiments. Because of the time-intensive nature of manual X-ray crystallographic refinement, automated refinement methods that thoroughly explore conformational space are essential for the systematic construction of structurally variable models. Using five proteins spanning resolutions of 1.0-2.8 A, it is demonstrated how torsion-angle sampling of backbone and side-chain libraries with filtering against both the chemical energy, using a modern effective potential, and the electron density, coupled with minimization of a reciprocal-space X-ray target function, can generate multiple structurally variable models which fit the X-ray data well. Torsion-angle sampling as implemented in the Protein Local Optimization Program (PLOP) has been used in this work. Models with the lowest R(free) values are obtained when electrostatic and implicit solvation terms are included in the effective potential. HIV-1 protease, calmodulin and SUMO-conjugating enzyme illustrate how variability in the ensemble of structures captures structural variability that is observed across multiple crystal structures and is linked to functional flexibility at hinge regions and binding interfaces. An ensemble-refinement procedure is proposed to differentiate between variability that is a consequence of physical conformational heterogeneity and that which reflects uncertainty in the atomic coordinates.
Figures

Similar articles
-
Exploring the dynamic information content of a protein NMR structure: comparison of a molecular dynamics simulation with the NMR and X-ray structures of Escherichia coli ribonuclease HI.Proteins. 1999 Jul 1;36(1):87-110. doi: 10.1002/(sici)1097-0134(19990701)36:1<87::aid-prot8>3.0.co;2-r. Proteins. 1999. PMID: 10373009
-
Conformational Flexibility of Ubiquitin-Modified and SUMO-Modified PCNA Shown by Full-Ensemble Hybrid Methods.J Mol Biol. 2018 Dec 7;430(24):5294-5303. doi: 10.1016/j.jmb.2018.10.017. Epub 2018 Oct 28. J Mol Biol. 2018. PMID: 30381149 Free PMC article.
-
Reintroducing electrostatics into protein X-ray structure refinement: bulk solvent treated as a dielectric continuum.Acta Crystallogr D Biol Crystallogr. 2003 Dec;59(Pt 12):2094-103. doi: 10.1107/s090744490301833x. Epub 2003 Nov 27. Acta Crystallogr D Biol Crystallogr. 2003. PMID: 14646067
-
Recent developments for the efficient crystallographic refinement of macromolecular structures.Curr Opin Struct Biol. 1998 Oct;8(5):606-11. doi: 10.1016/s0959-440x(98)80152-8. Curr Opin Struct Biol. 1998. PMID: 9818265 Review.
-
Combination of NMR spectroscopy and X-ray crystallography offers unique advantages for elucidation of the structural basis of protein complex assembly.Sci China Life Sci. 2011 Feb;54(2):101-11. doi: 10.1007/s11427-011-4137-2. Epub 2011 Feb 14. Sci China Life Sci. 2011. PMID: 21318479 Review.
Cited by
-
Automated identification of functional dynamic contact networks from X-ray crystallography.Nat Methods. 2013 Sep;10(9):896-902. doi: 10.1038/nmeth.2592. Epub 2013 Aug 4. Nat Methods. 2013. PMID: 23913260 Free PMC article.
-
Modeling discrete heterogeneity in X-ray diffraction data by fitting multi-conformers.Acta Crystallogr D Biol Crystallogr. 2009 Oct;65(Pt 10):1107-17. doi: 10.1107/S0907444909030613. Epub 2009 Sep 16. Acta Crystallogr D Biol Crystallogr. 2009. PMID: 19770508 Free PMC article.
-
Sparse estimation for structural variability.Algorithms Mol Biol. 2011 Apr 19;6:12. doi: 10.1186/1748-7188-6-12. Algorithms Mol Biol. 2011. PMID: 21504605 Free PMC article.
-
Protein flexibility: coordinate uncertainties and interpretation of structural differences.Acta Crystallogr D Biol Crystallogr. 2009 Nov;65(Pt 11):1140-61. doi: 10.1107/S090744490903145X. Epub 2009 Oct 22. Acta Crystallogr D Biol Crystallogr. 2009. PMID: 19923711 Free PMC article.
-
Multistart simulated annealing refinement of the crystal structure of the 70S ribosome.Proc Natl Acad Sci U S A. 2009 Oct 27;106(43):18195-200. doi: 10.1073/pnas.0909287106. Epub 2009 Oct 12. Proc Natl Acad Sci U S A. 2009. PMID: 19822758 Free PMC article.
References
-
- Alberts, B., Bray, D., Lewis, J., Raff, M., Roberts, K. & Watson, J. D. (2002). Molecular Biology of the Cell. New York: Garland Science.
-
- Andrec, M., Harano, Y., Jacobson, M. P., Friesner, R. A. & Levy, R. M. (2002). J. Struct. Funct. Genomics, 2, 103–111. - PubMed
-
- Bakker, P. I. de, DePristo, M. A., Burke, D. F. & Blundell, T. L. (2003). Proteins, 51, 21–40. - PubMed
-
- Berman, H., Henrick, K. & Nakamura, H. (2003). Nature Struct. Biol. 10, 980. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

