Characterization of the Fe site in iron-sulfur cluster-free hydrogenase (Hmd) and of a model compound via nuclear resonance vibrational spectroscopy (NRVS)
- PMID: 18407624
- PMCID: PMC2734977
- DOI: 10.1021/ic701251j
Characterization of the Fe site in iron-sulfur cluster-free hydrogenase (Hmd) and of a model compound via nuclear resonance vibrational spectroscopy (NRVS)
Abstract
We have used (57)Fe nuclear resonance vibrational spectroscopy (NRVS) to study the iron site in the iron-sulfur cluster-free hydrogenase Hmd from the methanogenic archaeon Methanothermobacter marburgensis. The spectra have been interpreted by comparison with a cis-(CO)2-ligated Fe model compound, Fe(S2C2H4)(CO)2(PMe3)2, as well as by normal mode simulations of plausible active site structures. For this model complex, normal mode analyses both from an optimized Urey-Bradley force field and from complementary density functional theory (DFT) calculations produced consistent results. For Hmd, previous IR spectroscopic studies found strong CO stretching modes at 1944 and 2011 cm(-1), interpreted as evidence for cis-Fe(CO)2 ligation. The NRVS data provide further insight into the dynamics of the Fe site, revealing Fe-CO stretch and Fe-CO bend modes at 494, 562, 590, and 648 cm(-1), consistent with the proposed cis-Fe(CO)2 ligation. The NRVS also reveals a band assigned to Fe-S stretching motion at approximately 311 cm(-1) and another reproducible feature at approximately 380 cm(-1). The (57)Fe partial vibrational densities of states (PVDOS) for Hmd can be reasonably well simulated by a normal mode analysis based on a Urey-Bradley force field for a five-coordinate cis-(CO)2-ligated Fe site with additional cysteine, water, and pyridone cofactor ligands. A "truncated" model without a water ligand can also be used to match the NRVS data. A final interpretation of the Hmd NRVS data, including DFT analysis, awaits a three-dimensional structure for the active site.
Figures

 ), and resonance Raman (
), and resonance Raman ( ) for Fe(S2C2H4)(CO)2(PMe3)2. Right: Molecular structure of Fe(S2C2H4)(CO)2(PMe3)2. Thermal ellipsoids are set at the 50% probability level. Hydrogen atoms are omitted for clarity. Key distances (Å) and angles (°): Fe(1)–C(1) 1.766(3), Fe(1)–C(2) 1.773(3), Fe(1)–P(2) 2.2628(9), Fe(1)–P(1) 2.2780(9), Fe(1)–S(2) 2.3181(11), Fe(1)-S(1) 2.3355(9); C(1)–Fe(1)–C(2) 97.56(13), C(1)–Fe(1)–P(2) 91.03(9), C(2)–Fe(1)–P(2) 92.22(9), C(1)–Fe(1)–P(1) 89.37(9), C(2)–Fe(1)–P(1) 87.27(9), P(2)–Fe(1)–P(1) 179.38(3), C(1)–Fe(1)–S(2) 174.70(10), C(2)–Fe(1)–S(2) 87.68(11), P(2)–Fe(1)–S(2) 87.92(3), P(1)–Fe(1)–S(2) 91.73(3), C(1)–Fe(1)–S(1) 86.22(9), C(2)–Fe(1)–S(1) 176.18(11), P(2)–Fe(1)–S(1) 87.17(4), P(1)–Fe(1)–S(1) 93.32(3), S(2)–Fe(1)–S(1) 88.53(3).
) for Fe(S2C2H4)(CO)2(PMe3)2. Right: Molecular structure of Fe(S2C2H4)(CO)2(PMe3)2. Thermal ellipsoids are set at the 50% probability level. Hydrogen atoms are omitted for clarity. Key distances (Å) and angles (°): Fe(1)–C(1) 1.766(3), Fe(1)–C(2) 1.773(3), Fe(1)–P(2) 2.2628(9), Fe(1)–P(1) 2.2780(9), Fe(1)–S(2) 2.3181(11), Fe(1)-S(1) 2.3355(9); C(1)–Fe(1)–C(2) 97.56(13), C(1)–Fe(1)–P(2) 91.03(9), C(2)–Fe(1)–P(2) 92.22(9), C(1)–Fe(1)–P(1) 89.37(9), C(2)–Fe(1)–P(1) 87.27(9), P(2)–Fe(1)–P(1) 179.38(3), C(1)–Fe(1)–S(2) 174.70(10), C(2)–Fe(1)–S(2) 87.68(11), P(2)–Fe(1)–S(2) 87.92(3), P(1)–Fe(1)–S(2) 91.73(3), C(1)–Fe(1)–S(1) 86.22(9), C(2)–Fe(1)–S(1) 176.18(11), P(2)–Fe(1)–S(1) 87.17(4), P(1)–Fe(1)–S(1) 93.32(3), S(2)–Fe(1)–S(1) 88.53(3).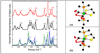
 ), Vibratz simulation (—), and DFT simulations using LACV3P**+ basis and PWPW91 functional (
), Vibratz simulation (—), and DFT simulations using LACV3P**+ basis and PWPW91 functional ( ) or B3LYP functional (– –) for Fe(S2C2H4)(CO)2(PMe3)2. Right: atomic motion in normal modes at (a) 626 cm−1 in Vibratz simulation, (b) 636 cm−1 in DFT simulation using PWPW91 functional and LACV3P**+ basis.
) or B3LYP functional (– –) for Fe(S2C2H4)(CO)2(PMe3)2. Right: atomic motion in normal modes at (a) 626 cm−1 in Vibratz simulation, (b) 636 cm−1 in DFT simulation using PWPW91 functional and LACV3P**+ basis.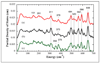
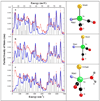
 ) in natural abundance water and simulations (
) in natural abundance water and simulations ( ) for (top to bottom): ‘tetrahedral’ model, 4-coordinate ‘truncated’ model, 5-coordinate ‘trigonal bipyramidal’ model with 16O water ligand. Right: Ball and stick models of the structures employed in the calculations. Additional cysteine and pyridone atoms have been omitted for clarity.
) for (top to bottom): ‘tetrahedral’ model, 4-coordinate ‘truncated’ model, 5-coordinate ‘trigonal bipyramidal’ model with 16O water ligand. Right: Ball and stick models of the structures employed in the calculations. Additional cysteine and pyridone atoms have been omitted for clarity.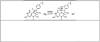
References
-
- Zirngibl C, Hedderich R, Thauer RK. N5, N10-Methylenetetrahydromethanopterin dehydrogenase from Methanobacterium thermoautotrophicum has hydrogenase activity. FEBS Lett. 1990;261:112–116.
-
- Zirngibl C, van Dongen W, Schwörer B, von Bünau R, Richter M, Klein A, Thauer RK. H2-forming methylenetetrahydromethanopterin dehydrogenase, a novel type of hydrogenase without iron-sulfur clusters in methanogenic archaea. Eur. J. Biochem. 1992;208:511–520. - PubMed
-
- Schleucher J, Griesinger C, Schwörer B, Thauer RK. H2-Forming N5,N10-Methylenetetrahydromethanopterin Dehydrogenase from Methanobacterium thermoautotrophicum Catalyzes a Stereoselective Hydride Transfer As Determined by Two-Dimensional NMR Spectroscopy. Biochem. 1994;33:3986–3983. - PubMed
-
- Thauer RK, Klein AR, Hartmann GC. Reactions with Molecular Hydrogen in Microorganisms: Evidence for a Purely Organic Hydrogenation Catalyst. Chem. Rev. 1996;97:3031–3042. - PubMed
-
- Shima S, Thauer RK. A third type of hydrogenase catalyzing H2 activation. Chem. Rec. 2007;7:37–46. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

