Dynamics of cellular focal adhesions on deformable substrates: consequences for cell force microscopy
- PMID: 18408038
- PMCID: PMC2440452
- DOI: 10.1529/biophysj.107.127399
Dynamics of cellular focal adhesions on deformable substrates: consequences for cell force microscopy
Abstract
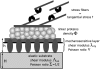
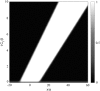
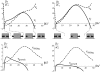
Cell focal adhesions are micrometer-sized aggregates of proteins that anchor the cell to the extracellular matrix. Within the cell, these adhesions are connected to the contractile, actin cytoskeleton; this allows the adhesions to transmit forces to the surrounding matrix and makes the adhesion assembly sensitive to the rigidity of their environment. In this article, we predict the dynamics of focal adhesions as a function of the rigidity of the substrate. We generalize previous theories and include the fact that the dynamics of proteins that adsorb to adhesions are also driven by their coupling to cell contractility and the deformation of the matrix. We predict that adhesions reach a finite size that is proportional to the elastic compliance of the substrate, on a timescale that also scales with the compliance: focal adhesions quickly reach a relatively small, steady-state size on soft materials. However, their apparent sliding is not sensitive to the rigidity of the substrate. We also suggest some experimental probes of these ideas and discuss the nature of information that can be extracted from cell force microscopy on deformable substrates.
Figures


Similar articles
-
Force-induced adsorption and anisotropic growth of focal adhesions.Biophys J. 2006 May 15;90(10):3469-84. doi: 10.1529/biophysj.105.074377. Epub 2006 Mar 2. Biophys J. 2006. PMID: 16513789 Free PMC article.
-
Limitation of cell adhesion by the elasticity of the extracellular matrix.Biophys J. 2006 Jul 1;91(1):61-73. doi: 10.1529/biophysj.105.077115. Epub 2006 Mar 31. Biophys J. 2006. PMID: 16581840 Free PMC article.
-
Cell mechanosensitivity controls the anisotropy of focal adhesions.Proc Natl Acad Sci U S A. 2004 Aug 24;101(34):12520-5. doi: 10.1073/pnas.0403539101. Epub 2004 Aug 16. Proc Natl Acad Sci U S A. 2004. PMID: 15314229 Free PMC article.
-
Traction force microscopy for understanding cellular mechanotransduction.BMB Rep. 2020 Feb;53(2):74-81. doi: 10.5483/BMBRep.2020.53.2.308. BMB Rep. 2020. PMID: 31964473 Free PMC article. Review.
-
Adhesion-dependent cell mechanosensitivity.Annu Rev Cell Dev Biol. 2003;19:677-95. doi: 10.1146/annurev.cellbio.19.111301.153011. Annu Rev Cell Dev Biol. 2003. PMID: 14570586 Review.
Cited by
-
A Review of Cell Adhesion Studies for Biomedical and Biological Applications.Int J Mol Sci. 2015 Aug 5;16(8):18149-84. doi: 10.3390/ijms160818149. Int J Mol Sci. 2015. PMID: 26251901 Free PMC article. Review.
-
Whole cell mechanics of contractile fibroblasts: relations between effective cellular and extracellular matrix moduli.Philos Trans A Math Phys Eng Sci. 2010 Feb 13;368(1912):635-54. doi: 10.1098/rsta.2009.0240. Philos Trans A Math Phys Eng Sci. 2010. PMID: 20047943 Free PMC article. Review.
-
Adhesion dynamics and durotaxis in migrating cells.Phys Biol. 2011 Feb;8(1):015011. doi: 10.1088/1478-3975/8/1/015011. Epub 2011 Feb 7. Phys Biol. 2011. PMID: 21301061 Free PMC article.
-
The mechanical microenvironment regulates ovarian cancer cell morphology, migration, and spheroid disaggregation.Sci Rep. 2018 May 8;8(1):7228. doi: 10.1038/s41598-018-25589-0. Sci Rep. 2018. PMID: 29740072 Free PMC article.
-
Excitable waves at the margin of the contact area between a cell and a substrate.Phys Biol. 2009 Jul 1;6(2):025010. doi: 10.1088/1478-3975/6/2/025010. Phys Biol. 2009. PMID: 19571372 Free PMC article.
References
-
- Engler, A. J., S. Sen, H. L. Sweeney, and D. E. Discher. 2006. Matrix elasticity directs stem cell lineage specification. Cell. 126:677–689. - PubMed
-
- Balaban, N. Q., U. S. Schwarz, D. Riveline, P. Goichberg, G. Tzur, I. Sabanay, D. Mahalu, S. A. Safran, A. Bershadsky, L. Addadi, and B. Geiger. 2001. Force and focal adhesion assembly: a close relationship studied using elastic micropatterned substrates. Nat. Cell Biol. 3:466–472. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

