Engineering BioBrick vectors from BioBrick parts
- PMID: 18410688
- PMCID: PMC2373286
- DOI: 10.1186/1754-1611-2-5
Engineering BioBrick vectors from BioBrick parts
Abstract
Background: The underlying goal of synthetic biology is to make the process of engineering biological systems easier. Recent work has focused on defining and developing standard biological parts. The technical standard that has gained the most traction in the synthetic biology community is the BioBrick standard for physical composition of genetic parts. Parts that conform to the BioBrick assembly standard are BioBrick standard biological parts. To date, over 2,000 BioBrick parts have been contributed to, and are available from, the Registry of Standard Biological Parts.
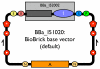
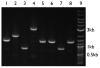
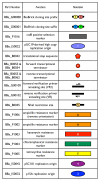
Results: Here we extended the same advantages of BioBrick standard biological parts to the plasmid-based vectors that are used to provide and propagate BioBrick parts. We developed a process for engineering BioBrick vectors from BioBrick parts. We designed a new set of BioBrick parts that encode many useful vector functions. We combined the new parts to make a BioBrick base vector that facilitates BioBrick vector construction. We demonstrated the utility of the process by constructing seven new BioBrick vectors. We also successfully used the resulting vectors to assemble and propagate other BioBrick standard biological parts.
Conclusion: We extended the principles of part reuse and standardization to BioBrick vectors. As a result, myriad new BioBrick vectors can be readily produced from all existing and newly designed BioBrick parts. We invite the synthetic biology community to (1) use the process to make and share new BioBrick vectors; (2) expand the current collection of BioBrick vector parts; and (3) characterize and improve the available collection of BioBrick vector parts.
Figures


References
-
- Weiss R. Cellular Computation and Communications using Engineered Genetic Regulatory Networks. PhD thesis, Massachusetts Institute of Technology, Department of Electrical Engineering and Computer Science. 2001. http://hdl.handle.net/1721.1/8228
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

