Accelerating string set matching in FPGA hardware for bioinformatics research
- PMID: 18412963
- PMCID: PMC2374783
- DOI: 10.1186/1471-2105-9-197
Accelerating string set matching in FPGA hardware for bioinformatics research
Abstract
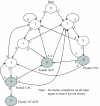
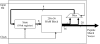
Background: This paper describes techniques for accelerating the performance of the string set matching problem with particular emphasis on applications in computational proteomics. The process of matching peptide sequences against a genome translated in six reading frames is part of a proteogenomic mapping pipeline that is used as a case-study. The Aho-Corasick algorithm is adapted for execution in field programmable gate array (FPGA) devices in a manner that optimizes space and performance. In this approach, the traditional Aho-Corasick finite state machine (FSM) is split into smaller FSMs, operating in parallel, each of which matches up to 20 peptides in the input translated genome. Each of the smaller FSMs is further divided into five simpler FSMs such that each simple FSM operates on a single bit position in the input (five bits are sufficient for representing all amino acids and special symbols in protein sequences).
Results: This bit-split organization of the Aho-Corasick implementation enables efficient utilization of the limited random access memory (RAM) resources available in typical FPGAs. The use of on-chip RAM as opposed to FPGA logic resources for FSM implementation also enables rapid reconfiguration of the FPGA without the place and routing delays associated with complex digital designs.
Conclusion: Experimental results show storage efficiencies of over 80% for several data sets. Furthermore, the FPGA implementation executing at 100 MHz is nearly 20 times faster than an implementation of the traditional Aho-Corasick algorithm executing on a 2.67 GHz workstation.
Figures
Similar articles
-
Hardware acceleration of processing of mass spectrometric data for proteomics.Bioinformatics. 2007 Mar 15;23(6):724-31. doi: 10.1093/bioinformatics/btl656. Epub 2007 Feb 3. Bioinformatics. 2007. PMID: 17277335
-
Hardware-accelerated protein identification for mass spectrometry.Rapid Commun Mass Spectrom. 2005;19(6):833-7. doi: 10.1002/rcm.1853. Rapid Commun Mass Spectrom. 2005. PMID: 15723443
-
Design of FPGA-Based SHE and SPWM Digital Switching Controllers for 21-Level Cascaded H-Bridge Multilevel Inverter Model.Micromachines (Basel). 2022 Jan 25;13(2):179. doi: 10.3390/mi13020179. Micromachines (Basel). 2022. PMID: 35208304 Free PMC article.
-
Machine learning algorithms for FPGA Implementation in biomedical engineering applications: A review.Heliyon. 2024 Feb 18;10(4):e26652. doi: 10.1016/j.heliyon.2024.e26652. eCollection 2024 Feb 29. Heliyon. 2024. PMID: 38434008 Free PMC article. Review.
-
The molecular scanner: concept and developments.Curr Opin Biotechnol. 2004 Feb;15(1):17-23. doi: 10.1016/j.copbio.2003.12.003. Curr Opin Biotechnol. 2004. PMID: 15102461 Review.
Cited by
-
A quick guide for developing effective bioinformatics programming skills.PLoS Comput Biol. 2009 Dec;5(12):e1000589. doi: 10.1371/journal.pcbi.1000589. Epub 2009 Dec 24. PLoS Comput Biol. 2009. PMID: 20041221 Free PMC article. No abstract available.
-
The proteogenomic mapping tool.BMC Bioinformatics. 2011 Apr 22;12:115. doi: 10.1186/1471-2105-12-115. BMC Bioinformatics. 2011. PMID: 21513508 Free PMC article.
-
Robust control of a wind energy conversion system: FPGA real-time implementation.Heliyon. 2024 Aug 3;10(15):e35712. doi: 10.1016/j.heliyon.2024.e35712. eCollection 2024 Aug 15. Heliyon. 2024. PMID: 39170361 Free PMC article.
References
-
- Jaffe JD, Berg HC, Church GM. Proteogenomic mapping as a complementary method to perform genome annotation. Proteomics. 2004;4:59–77. - PubMed
-
- Jaffe JD, Stange-Thomann N, Smith C, DeCaprio D, Fisher S, Butler J, Calvo S, Elkins T, FitzGerald MG, Hafez N, Kodira CD, Major J, Wang S, Wilkinson J, Nicol R, Nusbaum C, Birren B, Berg HC, Church GM. The complete genome and proteome of Mycoplasma mobile. Genome Res. 2004;14:1447–1461. - PMC - PubMed
-
- Kuster B, Mortensen P, Andersen JS, Mann M. Mass spectrometry allows direct identification of proteins in large genomes. Proteomics. 2001;1:641–650. - PubMed
-
- McCarthy FM, Cooksey AM, Wang N, Bridges SM, Pharr GT, Burgess SC. Modeling a whole organ using proteomics: the avian bursa of Fabricius. Proteomics. 2006;6:2759–2771. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources