Crystal structures of barley thioredoxin h isoforms HvTrxh1 and HvTrxh2 reveal features involved in protein recognition and possibly in discriminating the isoform specificity
- PMID: 18424513
- PMCID: PMC2386739
- DOI: 10.1110/ps.083460308
Crystal structures of barley thioredoxin h isoforms HvTrxh1 and HvTrxh2 reveal features involved in protein recognition and possibly in discriminating the isoform specificity
Abstract
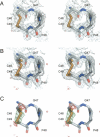
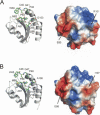
H-type thioredoxins (Trxs) constitute a particularly large Trx sub-group in higher plants. Here, the crystal structures are determined for the two barley Trx h isoforms, HvTrxh1 and HvTrxh2, in the partially radiation-reduced state to resolutions of 1.7 A, and for HvTrxh2 in the oxidized state to 2.0 A. The two Trxs have a sequence identity of 51% and highly similar fold and active-site architecture. Interestingly, the four independent molecules in the crystals of HvTrxh1 form two relatively large and essentially identical protein-protein interfaces. In each interface, a loop segment of one HvTrxh1 molecule is positioned along a shallow hydrophobic groove at the primary nucleophile Cys40 of another HvTrxh1 molecule. The association mode can serve as a model for the target protein recognition by Trx, as it brings the Met82 Cgamma atom (gamma position as a disulfide sulfur) of the bound loop segment in the proximity of the Cys40 thiol. The interaction involves three characteristic backbone-backbone hydrogen bonds in an antiparallel beta-sheet-like arrangement, similar to the arrangement observed in the structure of an engineered, covalently bound complex between Trx and a substrate protein, as reported by Maeda et al. in an earlier paper. The occurrence of an intermolecular salt bridge between Glu80 of the bound loop segment and Arg101 near the hydrophobic groove suggests that charge complementarity plays a role in the specificity of Trx. In HvTrxh2, isoleucine corresponds to this arginine, which emphasizes the potential for specificity differences between the coexisting barley Trx isoforms.
Figures



Similar articles
-
Kinetic and thermodynamic properties of two barley thioredoxin h isozymes, HvTrxh1 and HvTrxh2.FEBS Lett. 2010 Aug 4;584(15):3376-80. doi: 10.1016/j.febslet.2010.06.028. Epub 2010 Jun 23. FEBS Lett. 2010. PMID: 20594550
-
Identification, cloning and characterization of two thioredoxin h isoforms, HvTrxh1 and HvTrxh2, from the barley seed proteome.Eur J Biochem. 2003 Jun;270(12):2633-43. doi: 10.1046/j.1432-1033.2003.03637.x. Eur J Biochem. 2003. PMID: 12787030
-
The NADPH-dependent thioredoxin reductase/thioredoxin system in germinating barley seeds: gene expression, protein profiles, and interactions between isoforms of thioredoxin h and thioredoxin reductase.Plant Physiol. 2008 Feb;146(2):789-99. doi: 10.1104/pp.107.113639. Epub 2007 Dec 27. Plant Physiol. 2008. PMID: 18162587 Free PMC article.
-
Conferring specificity in redox pathways by enzymatic thiol/disulfide exchange reactions.Free Radic Res. 2016;50(2):206-45. doi: 10.3109/10715762.2015.1120864. Free Radic Res. 2016. PMID: 26573728 Review.
-
Thioredoxin and related proteins in procaryotes.FEMS Microbiol Rev. 1988 Dec;4(4):271-97. doi: 10.1111/j.1574-6968.1988.tb02747.x. FEMS Microbiol Rev. 1988. PMID: 3152490 Review.
Cited by
-
The conformational stability and biophysical properties of the eukaryotic thioredoxins of Pisum sativum are not family-conserved.PLoS One. 2011 Feb 22;6(2):e17068. doi: 10.1371/journal.pone.0017068. PLoS One. 2011. PMID: 21364950 Free PMC article.
-
Isolation, identification and sequence analysis of a thioredoxin h gene, a member of subgroup III of h-type Trxs from grape (Vitis vinifera L. cv. Askari).Mol Biol Rep. 2012 Apr;39(4):3683-93. doi: 10.1007/s11033-011-1143-1. Epub 2011 Jul 6. Mol Biol Rep. 2012. PMID: 21732058
-
Chaperone-like properties of tobacco plastid thioredoxins f and m.J Exp Bot. 2012 Jan;63(1):365-79. doi: 10.1093/jxb/err282. Epub 2011 Sep 23. J Exp Bot. 2012. PMID: 21948853 Free PMC article.
-
The specificity of thioredoxins and glutaredoxins is determined by electrostatic and geometric complementarity.Chem Sci. 2015 Dec 1;6(12):7049-7058. doi: 10.1039/c5sc01501d. Epub 2015 Sep 9. Chem Sci. 2015. PMID: 29861944 Free PMC article.
-
The barley grain thioredoxin system - an update.Front Plant Sci. 2013 May 21;4:151. doi: 10.3389/fpls.2013.00151. eCollection 2013. Front Plant Sci. 2013. PMID: 23734159 Free PMC article.
References
-
- Arnér, E.S.J., Holmgren, A. Physiological functions of thioredoxin and thioredoxin reductase. Eur. J. Biochem. 2000;267:6102–6109. - PubMed
-
- Bahadur, R.P., Chakrabarti, P., Rodier, F., Janin, J. Dissecting subunit interfaces in homodimeric proteins. Proteins. 2003;53:708–719. - PubMed
-
- Brünger, A.T., Adams, P.D., Clore, G.M., DeLano, W.L., Gros, P., Grosse-Kunstleve, R.W., Jiang, J.S., Kuszewski, J., Nilges, M., Pannu, N.S., et al. Crystallography and NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. 1998;D54:905–921. - PubMed
-
- Buchanan, B.B., Balmer, Y. Redox regulation: A broadening horizon. Annu. Rev. Plant Biol. 2005;56:187–220. - PubMed
-
- Capitani, G., Markovic-Housley, Z., DelVal, G., Morris, M., Jansonius, J.N., Schurmann, P. Crystal structures of two functionally different thioredoxins in spinach chloroplasts. J. Mol. Biol. 2000;302:135–154. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

