Minimal Escherichia coli cell for the most efficient production of ethanol from hexoses and pentoses
- PMID: 18424547
- PMCID: PMC2446564
- DOI: 10.1128/AEM.02708-07
Minimal Escherichia coli cell for the most efficient production of ethanol from hexoses and pentoses
Abstract
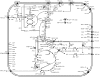
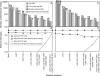
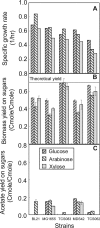
To obtain an efficient ethanologenic Escherichia coli strain, we reduced the functional space of the central metabolic network, with eight gene knockout mutations, from over 15,000 pathway possibilities to 6 pathway options that support cell function. The remaining pathways, identified by elementary mode analysis, consist of four pathways with non-growth-associated conversion of pentoses and hexoses into ethanol at theoretical yields and two pathways with tight coupling of anaerobic cell growth with ethanol formation at high yields. Elimination of three additional genes resulted in a strain that selectively grows only on pentoses, even in the presence of glucose, with a high ethanol yield. We showed that the ethanol yields of strains with minimized metabolic functionality closely matched the theoretical predictions. Remarkably, catabolite repression was completely absent during anaerobic growth, resulting in the simultaneous utilization of pentoses and hexoses for ethanol production.
Figures



References
-
- Alper, H., Y. Jin, J. F. Moxley, and G. Stephanopoulos. 2005. Identifying gene targets for the metabolic engineering of lycopene biosynthesis in Escherichia coli. Metab. Eng. 7:155-164. - PubMed
-
- Bachmann, B. J. 1996. Derivations and genotypes of some mutant derivatives of Escherichia coli K-12, p. 2460-2488. In F. C. Neidhardt, R. Curtiss III, J. L. Ingraham, E. C. C. Lin, K. B. Low, B. Magasanik, W. S. Reznikoff, M. Riley, M. Schaechter, and H. E. Umbarger (ed.), Escherichia coli and Salmonella: cellular and molecular biology, 2nd ed., vol. 2. ASM Press, Washington, DC.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

