A systematic survey in Arabidopsis thaliana of transcription factors that modulate circadian parameters
- PMID: 18426557
- PMCID: PMC2410138
- DOI: 10.1186/1471-2164-9-182
A systematic survey in Arabidopsis thaliana of transcription factors that modulate circadian parameters
Abstract
Background: Plant circadian systems regulate various biological processes in harmony with daily environmental changes. In Arabidopsis thaliana, the underlying clock mechanism is comprised of multiple integrated transcriptional feedbacks, which collectively lead to global patterns of rhythmic gene expression. The transcriptional networks are essential within the clock itself and in its output pathway.
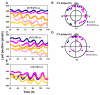
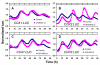
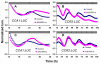
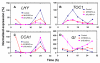
Results: Here, to expand understanding of transcriptional networks within and associated to the clock, we performed both an in silico analysis of transcript rhythmicity of transcription factor genes, and a pilot assessment of functional phenomics on the MYB, bHLH, and bZIP families. In our in silico analysis, we defined which members of these families express a circadian waveform of transcript abundance. Up to 20% of these families were over-represented as clock-controlled genes. To detect members that contribute to proper oscillator function, we systematically measured rhythmic growth via an imaging system in hundreds of misexpression lines targeting members of the transcription-factor families. Three transcription factors were found that conferred aberrant circadian rhythms when misexpressed: MYB3R2, bHLH69, and bHLH92.
Conclusion: Transcript abundance of many transcription factors in Arabidopsis oscillates in a circadian manner. Further, a developed pipeline assessed phenotypic contribution of a panel of transcriptional regulators in the circadian system.
Figures

Similar articles
-
From a repressilator-based circadian clock mechanism to an external coincidence model responsible for photoperiod and temperature control of plant architecture in Arabodopsis thaliana.Biosci Biotechnol Biochem. 2013;77(1):10-6. doi: 10.1271/bbb.120765. Epub 2013 Jan 7. Biosci Biotechnol Biochem. 2013. PMID: 23291766 Review.
-
Phytochrome-interacting factor 4 and 5 (PIF4 and PIF5) activate the homeobox ATHB2 and auxin-inducible IAA29 genes in the coincidence mechanism underlying photoperiodic control of plant growth of Arabidopsis thaliana.Plant Cell Physiol. 2011 Aug;52(8):1315-29. doi: 10.1093/pcp/pcr076. Epub 2011 Jun 11. Plant Cell Physiol. 2011. PMID: 21666227
-
A morning-specific phytohormone gene expression program underlying rhythmic plant growth.PLoS Biol. 2008 Sep 16;6(9):e225. doi: 10.1371/journal.pbio.0060225. PLoS Biol. 2008. PMID: 18798691 Free PMC article.
-
The Transcriptional Network in the Arabidopsis Circadian Clock System.Genes (Basel). 2020 Oct 29;11(11):1284. doi: 10.3390/genes11111284. Genes (Basel). 2020. PMID: 33138078 Free PMC article. Review.
-
MYB transcription factors in the Arabidopsis circadian clock.J Exp Bot. 2002 Jul;53(374):1551-7. doi: 10.1093/jxb/erf027. J Exp Bot. 2002. PMID: 12096093 Review.
Cited by
-
Recent advances in computational modeling as a conduit to understand the plant circadian clock.F1000 Biol Rep. 2010 Jul 14;2:49. doi: 10.3410/B2-49. F1000 Biol Rep. 2010. PMID: 20948785 Free PMC article.
-
Modulation of copper deficiency responses by diurnal and circadian rhythms in Arabidopsis thaliana.J Exp Bot. 2016 Jan;67(1):391-403. doi: 10.1093/jxb/erv474. Epub 2015 Oct 29. J Exp Bot. 2016. PMID: 26516126 Free PMC article.
-
AKIN10 activity as a cellular link between metabolism and circadian-clock entrainment in Arabidopsis thaliana.Plant Signal Behav. 2018 Mar 4;13(3):e1411448. doi: 10.1080/15592324.2017.1411448. Epub 2018 Apr 3. Plant Signal Behav. 2018. PMID: 29231782 Free PMC article.
-
Overexpression of HLH4 Inhibits Cell Elongation and Anthocyanin Biosynthesis in Arabidopsis thaliana.Cells. 2022 Mar 24;11(7):1087. doi: 10.3390/cells11071087. Cells. 2022. PMID: 35406652 Free PMC article.
-
Insights into nitrogen metabolism in the wild and cultivated lettuce as revealed by transcriptome and weighted gene co-expression network analysis.Sci Rep. 2022 Jun 14;12(1):9852. doi: 10.1038/s41598-022-13954-z. Sci Rep. 2022. PMID: 35701518 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

