N-terminal sequence analysis of proteins and peptides
- PMID: 18429102
- PMCID: PMC2917096
- DOI: 10.1002/0471140864.ps1110s08
N-terminal sequence analysis of proteins and peptides
Abstract
Amino-terminal (N-terminal) sequence analysis is used to identify the order of amino acids of proteins or peptides, starting at their N-terminal end. This unit describes the sequence analysis of protein or peptide samples in solution or bound to PVDF membranes using a Perkin-Elmer Procise Sequencer. Sequence analysis of protein or peptide samples in solution or bound to PVDF membranes using a Hewlett-Packard Model G1005A sequencer is also described. Methods are provided for optimizing separation of PTH amino acid derivatives on Perkin-Elmer instruments and for increasing the proportion of sample injected onto the PTH analyzer on older Perkin-Elmer instruments by installing a modified sample loop. The amount of data obtained from a single sequencer run is substantial, and careful interpretation of this data by an experienced scientist familiar with the current operation performance of the instrument used for this analysis is critically important. A discussion of data interpretation is therefore provided. Finally, discussion of optimization of sequencer performance as well as possible solutions to frequently encountered problems is included.
Figures


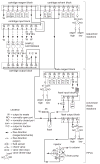


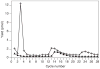
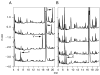
Similar articles
-
A method for high-performance sequence analysis using polyvinylidene difluoride membranes with a biphasic reaction column sequencer.Anal Biochem. 1994 Jan;216(1):213-22. doi: 10.1006/abio.1994.1027. Anal Biochem. 1994. PMID: 8135354
-
High-sensitivity gas phase sequence analysis of proteins and peptides on PVDF membranes using short cycle times.Anal Biochem. 1993 Oct;214(1):87-95. doi: 10.1006/abio.1993.1460. Anal Biochem. 1993. PMID: 8250259
-
Microsequencing of proteins and peptides in the Knauer sequencer with and without covalent attachment to polyvinylidene difluoride membranes by the wet-phase degradation technique.Biol Chem Hoppe Seyler. 1991 May;372(5):351-61. doi: 10.1515/bchm3.1991.372.1.351. Biol Chem Hoppe Seyler. 1991. PMID: 1872998
-
The chemistry of protein sequence analysis.EXS. 2000;88:99-117. doi: 10.1007/978-3-0348-8458-7_7. EXS. 2000. PMID: 10803374 Review.
-
A brief review of other notable protein blotting methods.Methods Mol Biol. 2009;536:367-84. doi: 10.1007/978-1-59745-542-8_38. Methods Mol Biol. 2009. PMID: 19378075 Review.
Cited by
-
PEDF improves cardiac function in rats subjected to myocardial ischemia/reperfusion injury by inhibiting ROS generation via PEDF‑R.Int J Mol Med. 2018 Jun;41(6):3243-3252. doi: 10.3892/ijmm.2018.3552. Epub 2018 Mar 9. Int J Mol Med. 2018. PMID: 29532859 Free PMC article.
-
L-Amino Acid Oxidases From Mushrooms Show Antibacterial Activity Against the Phytopathogen Ralstonia solanacearum.Front Microbiol. 2020 May 19;11:977. doi: 10.3389/fmicb.2020.00977. eCollection 2020. Front Microbiol. 2020. PMID: 32508788 Free PMC article.
-
Aldolases a and C are ribonucleolytic components of a neuronal complex that regulates the stability of the light-neurofilament mRNA.J Neurosci. 2005 Apr 27;25(17):4353-64. doi: 10.1523/JNEUROSCI.0885-05.2005. J Neurosci. 2005. PMID: 15858061 Free PMC article.
-
Rational design of an orthogonal tryptophanyl nonsense suppressor tRNA.Nucleic Acids Res. 2010 Oct;38(19):6813-30. doi: 10.1093/nar/gkq521. Epub 2010 Jun 22. Nucleic Acids Res. 2010. PMID: 20571084 Free PMC article.
References
-
- Atherton D, Fernandez J, DeMott M, Andrews L, Mische SM. Routine protein sequence analysis below ten picomoles: One sequencing facility’s approach. In: Angelletti R, editor. Techniques in Protein Chemistry IV. Academic Press; San Diego: 1993. pp. 409–418.
-
- Beyer K, Bardina L, Grishina G, Sampson HA. Identification of sesame seed allergens by 2-dimensional proteomics and Edman sequencing: see storage proteins as common food antigens. J Allergy Clin Immunol. 2002;110:154–159. - PubMed
-
- Brown JL, Roberts WK. Evidence that ~80% of the soluble proteins from Ehrlich ascites cells are N-alpha acetylated. J Biol Chem. 1976;251:1009–1014. - PubMed
-
- Edman P. A method for the determination of the amino acid sequence in peptides. Arch Biochem Biophys. 1949;22:475–480. - PubMed
-
- Edman P, Begg G. A protein sequenator. Eur J Biochem. 1967;1:80–91. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

