Structural analysis of human immunodeficiency virus type 1 CRF01_AE protease in complex with the substrate p1-p6
- PMID: 18434392
- PMCID: PMC2447101
- DOI: 10.1128/JVI.00018-08
Structural analysis of human immunodeficiency virus type 1 CRF01_AE protease in complex with the substrate p1-p6
Abstract
The effect of amino acid variability between human immunodeficiency virus type 1 (HIV-1) clades on structure and the emergence of resistance mutations in HIV-1 protease has become an area of significant interest in recent years. We determined the first crystal structure of the HIV-1 CRF01_AE protease in complex with the p1-p6 substrate to a resolution of 2.8 A. Hydrogen bonding between the flap hinge and the protease core regions shows significant structural rearrangements in CRF01_AE protease compared to the clade B protease structure.
Figures
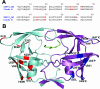

References
-
- Ariyoshi, K., M. Matsuda, H. Miura, S. Tateishi, K. Yamada, and W. Sugiura. 2003. Patterns of point mutations associated with antiretroviral drug treatment failure in CRF01_AE (subtype E) infection differ from subtype B infection. J. Acquir. Immune Defic. Syndr. 33336-342. - PubMed
-
- Becker-Pergola, G., P. Kataaha, L. Johnston-Dow, S. Fung, J. B. Jackson, and S. H. Eshleman. 2000. Analysis of HIV type 1 protease and reverse transcriptase in antiretroviral drug-naive Ugandan adults. AIDS Res. Hum. Retrovir. 16807-813. - PubMed
-
- Clemente, J. C., R. M. Coman, M. M. Thiaville, L. K. Janka, J. A. Jeung, S. Nukoolkarn, L. Govindasamy, M. Agbandje-McKenna, R. McKenna, W. Leelamanit, M. M. Goodenow, and B. M. Dunn. 2006. Analysis of HIV-1 CRF_01 A/E protease inhibitor resistance: structural determinants for maintaining sensitivity and developing resistance to atazanavir. Biochemistry 455468-5477. - PMC - PubMed
-
- Emsley, P., and K. Cowtan. 2004. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 602126-2132. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

