Toward a comprehensive temperature-sensitive mutant repository of the essential genes of Saccharomyces cerevisiae
- PMID: 18439903
- PMCID: PMC4130347
- DOI: 10.1016/j.molcel.2008.02.021
Toward a comprehensive temperature-sensitive mutant repository of the essential genes of Saccharomyces cerevisiae
Abstract
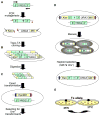
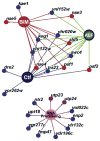
The Saccharomyces cerevisiae gene deletion project revealed that approximately 20% of yeast genes are required for viability. The analysis of essential genes traditionally relies on conditional mutants, typically temperature-sensitive (ts) alleles. We developed a systematic approach (termed "diploid shuffle") useful for generating a ts allele for each essential gene in S. cerevisiae and for improved genetic manipulation of mutant alleles and gene constructs in general. Importantly, each ts allele resides at its normal genomic locus, flanked by specific cognate UPTAG and DNTAG bar codes. A subset of 250 ts mutants, including ts alleles for all uncharacterized essential genes and prioritized for genes with human counterparts, is now ready for distribution. The importance of this collection is demonstrated by biochemical and genetic screens that reveal essential genes involved in RNA processing and maintenance of chromosomal stability.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

