REBELOTE, SQUINT, and ULTRAPETALA1 function redundantly in the temporal regulation of floral meristem termination in Arabidopsis thaliana
- PMID: 18441215
- PMCID: PMC2390727
- DOI: 10.1105/tpc.107.053306
REBELOTE, SQUINT, and ULTRAPETALA1 function redundantly in the temporal regulation of floral meristem termination in Arabidopsis thaliana
Abstract
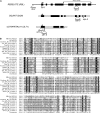
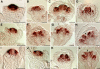
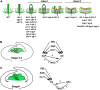
In Arabidopsis thaliana, flowers are determinate, showing a fixed number of whorls. Here, we report on three independent genes, a novel gene REBELOTE (RBL; protein of unknown function), SQUINT (SQN; a cyclophilin), and ULTRAPETALA1 (ULT1; a putative transcription factor) that redundantly influence floral meristem (FM) termination. Their mutations, combined with each other or with crabs claw, the genetic background in which they were isolated, trigger a strong FM indeterminacy with reiterations of extra floral whorls in the center of the flower. The range of phenotypes suggests that, in Arabidopsis, FM termination is initiated from stages 3 to 4 onwards and needs to be maintained through stage 6 and beyond, and that RBL, SQN, and ULT1 are required for this continuous regulation. We show that mutant phenotypes result from a decrease of AGAMOUS (AG) expression in an inner 4th whorl subdomain. However, the defect of AG activity alone does not explain all reported phenotypes, and our genetic data suggest that RBL, SQN, and, to a lesser extent, ULT1 also influence SUPERMAN activity. Finally, from all the molecular and genetic data presented, we argue that these genes contribute to the more stable and uniform development of flowers, termed floral developmental homeostasis.
Figures







References
-
- Alvarez, J., and Smyth, D.R. (1999). CRABS CLAW and SPATULA, two Arabidopsis genes that control carpel development in parallel with AGAMOUS. Development 126 2377–2386. - PubMed
-
- Angenent, G.C., Stuurman, J., Snowden, K.C., and Koes, R. (2005). Use of Petunia to unravel plant meristem functioning. Trends Plant Sci. 10 243–250. - PubMed
-
- Baum, D.A., and Hileman, L.C. (2006). A genetic model for the origin of flowers. In Flowering and Its Manipulation, C. Ainsworth, ed (Sheffield, UK: Blackwell Publishing), pp. 3–27.
-
- Bechtold, N., and Pelletier, G. (1998). In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. Methods Mol. Biol. 82 259–266. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

