Transcriptome analysis identifies pathways associated with enhanced maternal performance in QSi5 mice
- PMID: 18442417
- PMCID: PMC2386486
- DOI: 10.1186/1471-2164-9-197
Transcriptome analysis identifies pathways associated with enhanced maternal performance in QSi5 mice
Abstract
Background: Highly fecund mouse strains provide an ideal model to understand the factors affecting maternal performance. The QSi5 inbred strain of mice was selected for high fecundity and low inter-litter interval, and is very successful at weaning large numbers of offspring when compared to other inbred strains.
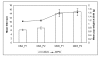
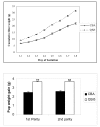
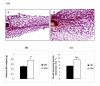
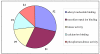
Results: Post-natal pup weight gain was used to estimate mammary gland output and to compare the performance of QSi5 mice to CBA mice. Cumulative litter weights and individual pup weight gain was significantly higher throughout the first eight days of lactation in QSi5 mice compared to CBA mice. Morphometric analysis of mammary glands during pregnancy in QSi5 mice revealed a 150 percent greater ductal side branching compared to CBA mice (P < 0.001). Ontology and pathway classification of transcript profiles from the two strains identified an enrichment of genes involved in a number of pathways, including the MAPK, tight junction, insulin signalling and Wnt signalling. Eleven of these genes, including six genes from the MAPK signalling pathway, were identified as associated with postnatal growth. Further, positive mediators of Wnt signalling, including Wnt4, Csnk2a1 and Smad4, were over-represented in the QSi5 strain profile, while negative regulators, including Dkkl1, Ppp2r1a and Nlk, were under-represented. These findings are consistent with the role of Wnt and MAPK signalling pathway in ductal morphogenesis and lobuloalveolar development suggesting enhanced activity in QSi5 mice. A similar pattern of phenotype concordance was seen amongst 12 genes from the tight junction pathway, but a pattern did not emerge from the insulin signalling genes. Amongst a group of differentially expressed imprinted genes, two maternal imprinted genes that suppress growth induced via the IGF signalling pathway, Grb10 and Igf2r, were under-represented in QSi5 mice. Whereas Peg3 and Plagl1, both paternally imprinted genes that enhance neonatal growth, were over-represented in QSi5 mice.
Conclusion: We propose that the combined action of at least three major signalling pathways involved in mammary gland development and milk secretion, namely Wnt, MAPK and tight junction pathways, contribute to the superior maternal performance phenotype in QSi5 mice. Additionally, favourable expression patterns of the imprinted genes Peg3, Plagl1, Grb10 and Igf2r may also contribute.
Figures


Similar articles
-
Identification of gene sets and pathways associated with lactation performance in mice.Physiol Genomics. 2013 Mar 1;45(5):171-81. doi: 10.1152/physiolgenomics.00139.2011. Epub 2013 Jan 2. Physiol Genomics. 2013. PMID: 23284081
-
Lactational performance of Quackenbush Swiss line 5 mice.J Anim Sci. 2006 Aug;84(8):2118-25. doi: 10.2527/jas.2005-609. J Anim Sci. 2006. PMID: 16864872
-
An integrative genomic analysis of the superior fecundity phenotype in QSi5 mice.Mol Biotechnol. 2013 Feb;53(2):217-26. doi: 10.1007/s12033-012-9530-y. Mol Biotechnol. 2013. PMID: 22447533
-
Wnt signalling in murine postnatal mammary gland development.Acta Physiol (Oxf). 2012 Jan;204(1):118-27. doi: 10.1111/j.1748-1716.2011.02283.x. Epub 2011 Apr 22. Acta Physiol (Oxf). 2012. PMID: 21518264 Review.
-
Control of the three-dimensional growth pattern of mammary epithelium: role of genes of the Wnt and erbB families studied using reconstituted epithelium.Biochem Soc Symp. 1998;63:21-34. Biochem Soc Symp. 1998. PMID: 9513708 Review.
Cited by
-
In silico QTL mapping of maternal nurturing ability with the mouse diversity panel.Physiol Genomics. 2012 Aug 17;44(16):787-98. doi: 10.1152/physiolgenomics.00159.2011. Epub 2012 Jul 3. Physiol Genomics. 2012. PMID: 22759921 Free PMC article.
-
Molecular genetics of adult ADHD: converging evidence from genome-wide association and extended pedigree linkage studies.J Neural Transm (Vienna). 2008 Nov;115(11):1573-85. doi: 10.1007/s00702-008-0119-3. Epub 2008 Oct 7. J Neural Transm (Vienna). 2008. PMID: 18839057
-
Sequencing the transcriptome of milk production: milk trumps mammary tissue.BMC Genomics. 2013 Dec 12;14:872. doi: 10.1186/1471-2164-14-872. BMC Genomics. 2013. PMID: 24330573 Free PMC article.
-
Fur removal promotes an earlier expression of involution-related genes in mammary gland of lactating mice.J Comp Physiol B. 2023 Mar;193(2):171-192. doi: 10.1007/s00360-023-01474-9. Epub 2023 Jan 18. J Comp Physiol B. 2023. PMID: 36650338 Free PMC article.
-
Characterization of conserved and nonconserved imprinted genes in swine.Biol Reprod. 2009 Nov;81(5):906-20. doi: 10.1095/biolreprod.109.078139. Epub 2009 Jul 1. Biol Reprod. 2009. PMID: 19571260 Free PMC article.
References
-
- Rudolph MC, McManaman J, Phang T, Russell T, Kominsky DJ, Serkova NJ, Stein T, Anderson SM, Neville MC. Metabolic regulation in the lactating mammary gland: A lipid synthesizing machine. Physiol Genomics. 2006;14:14. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

