Identification of novel tumor markers in prostate, colon and breast cancer by unbiased methylation profiling
- PMID: 18446232
- PMCID: PMC2323612
- DOI: 10.1371/journal.pone.0002079
Identification of novel tumor markers in prostate, colon and breast cancer by unbiased methylation profiling
Erratum in
- PLoS ONE. 2008;3(5). doi: 10.1371/annotation/2548989f-1f13-4ea5-8af8-62420b0a590e doi: 10.1371/annotation/2548989f-1f13-4ea5-8af8-62420b0a590e
Abstract
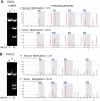
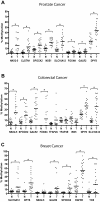
DNA hypermethylation is a common epigenetic abnormality in cancer and may serve as a useful marker to clone cancer-related genes as well as a marker of clinical disease activity. To identify CpG islands methylated in prostate cancer, we used methylated CpG island amplification (MCA) coupled with representational difference analysis (RDA) on prostate cancer cell lines. We isolated 34 clones that corresponded to promoter CpG islands, including 5 reported targets of hypermethylation in cancer. We confirmed the data for 17 CpG islands by COBRA and/or pyrosequencing. All 17 genes were methylated in at least 2 cell lines of a 21-cancer cell line panel containing prostate cancer, colon cancer, leukemia, and breast cancer. Based on methylation in primary tumors compared to normal adjacent tissues, NKX2-5, CLSTN1, SPOCK2, SLC16A12, DPYS and NSE1 are candidate biomarkers for prostate cancer (methylation range 50%-85%). The combination of NSE1 or SPOCK2 hypermethylation showed a sensitivity of 80% and specificity of 95% in differentiating cancer from normal. Similarly NKX2-5, SPOCK2, SLC16A12, DPYS and GALR2 are candidate biomarkers for colon cancer (methylation range 60%-95%) and GALR2 hypermethylation showed a sensitivity of 85% and specificity of 95%. Finally, SLC16A12, GALR2, TOX, SPOCK2, EGFR5 and DPYS are candidate biomarkers for breast cancer (methylation range 33%-79%) with the combination of EGFR5 or TOX hypermethylation showing a sensitivity of 92% and specificity of 92%. Expression analysis for eight genes that had the most hypermethylation confirmed the methylation associated silencing and reactivation with 5-aza-2'-deoxycytidine treatment. Our data identify new targets of transcriptional silencing in cancer, and provide new biomarkers that could be useful in screening for prostate cancer and other cancers.
Conflict of interest statement
Figures



References
-
- Jemal A, Murray T, Ward E, Samuels A, Tiwari RC, et al. Cancer statistics, 2005. Ca-A Cancer Journal for Clinicians. 2005;55:10–30. - PubMed
-
- Herman JG, Baylin SB. Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med. 2003;349:2042–2054. - PubMed
-
- Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JPJ. Alterations in DNA methylation - A fundamental aspect of neoplasia. Adv Cancer Res. 1998;72:141–196. - PubMed
-
- Kang GH, Lee S, Lee HJ, Hwang KS. Aberrant CpG island hypermethylation of multiple genes in prostate cancer and prostatic intraepithelial neoplasia. Journal of Pathology. 2004;202:233–240. - PubMed
-
- Yamanaka M, Watanabe M, Yamada Y, Takagi A, Murata T, et al. Altered methylation of multiple genes in carcinogenesis of the prostate. International Journal of Cancer. 2003;106:382–387. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

