Gene structure evolution of the Na+-Ca2+ exchanger (NCX) family
- PMID: 18447948
- PMCID: PMC2408596
- DOI: 10.1186/1471-2148-8-127
Gene structure evolution of the Na+-Ca2+ exchanger (NCX) family
Abstract
Background: The Na+-Ca2+ exchanger (NCX) is an important regulator of cytosolic Ca2+ levels. Many of its structural features are highly conserved across a wide range of species. Invertebrates have a single NCX gene, whereas vertebrate species have multiple NCX genes as a result of at least two duplication events. To examine the molecular evolution of NCX genes and understand the role of duplicated genes in the evolution of the vertebrate NCX gene family, we carried out phylogenetic analyses of NCX genes and compared NCX gene structures from sequenced genomes and individual clones.
Results: A single NCX in invertebrates and the protochordate Ciona, and the presence of at least four NCX genes in the genomes of teleosts, an amphibian, and a reptile suggest that a four member gene family arose in a basal vertebrate. Extensive examination of mammalian and avian genomes and synteny analysis argue that NCX4 may be lost in these lineages. Duplicates for NCX1, NCX2, and NCX4 were found in all sequenced teleost genomes. The presence of seven genes encoding NCX homologs may provide teleosts with the functional specialization analogous to the alternate splicing strategy seen with the three NCX mammalian homologs.
Conclusion: We have demonstrated that NCX4 is present in teleost, amphibian and reptilian species but has been secondarily and independently lost in mammals and birds. Comparative studies on conserved vertebrate homologs have provided a possible evolutionary route taken by gene duplicates subfunctionalization by minimizing homolog number.
Figures
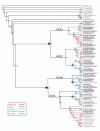
 ) indicate NCX duplication after Ciona intestinalis divergence and the four-point stars (
) indicate NCX duplication after Ciona intestinalis divergence and the four-point stars ( ) indicate teleost-specific NCX duplication. Branches are colour coded for vertebrate species, in which red designates mammals, purple indicates birds/aves, teal branches are for reptiles, green are amphibians, and blue are teleosts/fish. The dashed branch (---) indicates the division between invertebrate and vertebrate NCX sequences. The asterisk (*) indicates complete transcripts and those without an asterisk are genomic sequences. The letters 'a' and 'b' correspond to the teleost duplicate versions of NCX1, NCX2 and NCX4. Underlined species are new NCX sequences added in comparison to a previous paper [20].
) indicate teleost-specific NCX duplication. Branches are colour coded for vertebrate species, in which red designates mammals, purple indicates birds/aves, teal branches are for reptiles, green are amphibians, and blue are teleosts/fish. The dashed branch (---) indicates the division between invertebrate and vertebrate NCX sequences. The asterisk (*) indicates complete transcripts and those without an asterisk are genomic sequences. The letters 'a' and 'b' correspond to the teleost duplicate versions of NCX1, NCX2 and NCX4. Underlined species are new NCX sequences added in comparison to a previous paper [20].



Similar articles
-
Phylogeny of Na+/Ca2+ exchanger (NCX) genes from genomic data identifies new gene duplications and a new family member in fish species.Physiol Genomics. 2005 Apr 14;21(2):161-73. doi: 10.1152/physiolgenomics.00286.2004. Epub 2005 Mar 1. Physiol Genomics. 2005. PMID: 15741504
-
cDNA cloning and expression of the cardiac Na+/Ca2+ exchanger from Mozambique tilapia (Oreochromis mossambicus) reveal a teleost membrane transporter with mammalian temperature dependence.J Biol Chem. 2005 Aug 12;280(32):28903-11. doi: 10.1074/jbc.M504807200. Epub 2005 Jun 3. J Biol Chem. 2005. PMID: 15937330
-
Analysis of the Na+/Ca2+ exchanger gene family within the phylum Nematoda.PLoS One. 2014 Nov 14;9(11):e112841. doi: 10.1371/journal.pone.0112841. eCollection 2014. PLoS One. 2014. PMID: 25397810 Free PMC article.
-
Molecular biological studies of the cardiac sodium-calcium exchanger.Ann N Y Acad Sci. 1996 Apr 15;779:103-9. doi: 10.1111/j.1749-6632.1996.tb44774.x. Ann N Y Acad Sci. 1996. PMID: 8659815 Review.
-
Functional regulation of alternatively spliced Na+/Ca2+ exchanger (NCX1) isoforms.Ann N Y Acad Sci. 2002 Nov;976:187-96. doi: 10.1111/j.1749-6632.2002.tb04740.x. Ann N Y Acad Sci. 2002. PMID: 12502560 Review.
Cited by
-
Identification and apical membrane localization of an electrogenic Na⁺/Ca²⁺ exchanger NCX2a likely to be involved in renal Ca²⁺ excretion by seawater fish.Am J Physiol Regul Integr Comp Physiol. 2011 Nov;301(5):R1427-39. doi: 10.1152/ajpregu.00165.2011. Epub 2011 Aug 31. Am J Physiol Regul Integr Comp Physiol. 2011. PMID: 21880864 Free PMC article.
-
Cardiac contractility of the African sharptooth catfish, Clarias gariepinus: role of extracellular Ca2+, sarcoplasmic reticulum, and β-adrenergic stimulation.Fish Physiol Biochem. 2021 Dec;47(6):1969-1982. doi: 10.1007/s10695-021-01023-7. Epub 2021 Oct 19. Fish Physiol Biochem. 2021. PMID: 34668117
-
Ion Channel and Transporter Involvement in Chemotherapy-Induced Peripheral Neurotoxicity.Int J Mol Sci. 2024 Jun 14;25(12):6552. doi: 10.3390/ijms25126552. Int J Mol Sci. 2024. PMID: 38928257 Free PMC article. Review.
-
Evolutionary genomics reveals lineage-specific gene loss and rapid evolution of a sperm-specific ion channel complex: CatSpers and CatSperbeta.PLoS One. 2008;3(10):e3569. doi: 10.1371/journal.pone.0003569. Epub 2008 Oct 30. PLoS One. 2008. PMID: 18974790 Free PMC article.
-
Sodium-Calcium Exchangers of the SLC8 Family in Oligodendrocytes: Functional Properties in Health and Disease.Neurochem Res. 2020 Jun;45(6):1287-1297. doi: 10.1007/s11064-019-02949-4. Epub 2020 Jan 11. Neurochem Res. 2020. PMID: 31927687 Free PMC article. Review.
References
-
- Bers DM. Ca2+ influx and sarcoplasmic reticulum Ca2+ release in cardiac muscle activation during postrest recovery. Am J Physiol Heart Circ Physiol. 1985;248(3 Pt 2):H366–H381. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

