NOBAI: a web server for character coding of geometrical and statistical features in RNA structure
- PMID: 18448469
- PMCID: PMC2447726
- DOI: 10.1093/nar/gkn220
NOBAI: a web server for character coding of geometrical and statistical features in RNA structure
Abstract
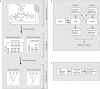
The Numeration of Objects in Biology: Alignment Inferences (NOBAI) web server provides a web interface to the applications in the NOBAI software package. This software codes topological and thermodynamic information related to the secondary structure of RNA molecules as multi-state phylogenetic characters, builds character matrices directly in NEXUS format and provides sequence randomization options. The web server is an effective tool that facilitates the search for evolutionary history embedded in the structure of functional RNA molecules. The NOBAI web server is accessible at 'http://www.manet.uiuc.edu/nobai/nobai.php'. This web site is free and open to all users and there is no login requirement.
Figures

References
-
- Hermann T, Patel DJ. Stitching together RNA tertiary structures. J. Mol. Biol. 1999;294:829–849. - PubMed
-
- Billoud B, Guerrucci MA, Masselot M, Deutsch JS. Cirripede phylogeny using a novel approach: molecular morphometrics. Mol. Biol. Evol. 2000;17:1435–1445. - PubMed
-
- Collins LJ, Moulton V, Penny D. Use of RNA secondary structure for studying the evolution of RNase P and RNase MRP. J. Mol. Evol. 2000;51:194–2004. - PubMed
-
- Caetano-Anollés G. Novel strategies to study the role of mutation and nucleic acid structure in evolution. Plant Cell Tissue Organ Cult. 2001;67:115–132.

