Access to ribosomal protein Rpl25p by the signal recognition particle is required for efficient cotranslational translocation
- PMID: 18448667
- PMCID: PMC2441686
- DOI: 10.1091/mbc.e07-10-1074
Access to ribosomal protein Rpl25p by the signal recognition particle is required for efficient cotranslational translocation
Abstract
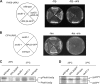
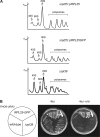
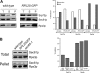
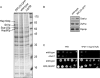
Targeting of proteins to the endoplasmic reticulum (ER) occurs cotranslationally necessitating the interaction of the signal recognition particle (SRP) and the translocon with the ribosome. Biochemical and structural studies implicate ribosomal protein Rpl25p as a major ribosome interaction site for both these factors. Here we characterize an RPL25GFP fusion, which behaves as a dominant mutant leading to defects in co- but not posttranslational translocation in vivo. In these cells, ribosomes still interact with ER membrane and the translocon, but are defective in binding SRP. Overexpression of SRP can restore ribosome binding of SRP, but only partially rescues growth and translocation defects. Our results indicate that Rpl25p plays a critical role in the recruitment of SRP to the ribosome.
Figures


Similar articles
-
Cotranslational signal-independent SRP preloading during membrane targeting.Nature. 2016 Aug 11;536(7615):224-8. doi: 10.1038/nature19309. Epub 2016 Aug 3. Nature. 2016. PMID: 27487213 Free PMC article.
-
Binding of signal recognition particle gives ribosome/nascent chain complexes a competitive advantage in endoplasmic reticulum membrane interaction.Mol Biol Cell. 1998 Jan;9(1):103-15. doi: 10.1091/mbc.9.1.103. Mol Biol Cell. 1998. PMID: 9436994 Free PMC article.
-
NAC functions as a modulator of SRP during the early steps of protein targeting to the endoplasmic reticulum.Mol Biol Cell. 2012 Aug;23(16):3027-40. doi: 10.1091/mbc.E12-02-0112. Epub 2012 Jun 27. Mol Biol Cell. 2012. PMID: 22740632 Free PMC article.
-
Translocation of proteins across the endoplasmic reticulum membrane.Int Rev Cytol. 1998;178:277-328. doi: 10.1016/s0074-7696(08)62139-7. Int Rev Cytol. 1998. PMID: 9348672 Review.
-
Co-translational targeting and translocation of proteins to the endoplasmic reticulum.Biochim Biophys Acta. 2013 Nov;1833(11):2392-402. doi: 10.1016/j.bbamcr.2013.02.021. Epub 2013 Feb 26. Biochim Biophys Acta. 2013. PMID: 23481039 Review.
Cited by
-
Co-translational localization of an LTR-retrotransposon RNA to the endoplasmic reticulum nucleates virus-like particle assembly sites.PLoS Genet. 2014 Mar 6;10(3):e1004219. doi: 10.1371/journal.pgen.1004219. eCollection 2014 Mar. PLoS Genet. 2014. PMID: 24603646 Free PMC article.
-
Defining the specificity of cotranslationally acting chaperones by systematic analysis of mRNAs associated with ribosome-nascent chain complexes.PLoS Biol. 2011 Jul;9(7):e1001100. doi: 10.1371/journal.pbio.1001100. Epub 2011 Jul 12. PLoS Biol. 2011. PMID: 21765803 Free PMC article.
-
The Not4 RING E3 Ligase: A Relevant Player in Cotranslational Quality Control.ISRN Mol Biol. 2013 Jan 21;2013:548359. doi: 10.1155/2013/548359. eCollection 2013. ISRN Mol Biol. 2013. PMID: 27335678 Free PMC article. Review.
-
The translational regulatory function of SecM requires the precise timing of membrane targeting.Mol Microbiol. 2011 Jul;81(2):540-53. doi: 10.1111/j.1365-2958.2011.07713.x. Epub 2011 Jun 3. Mol Microbiol. 2011. PMID: 21635582 Free PMC article.
-
Targeting of Proteins for Translocation at the Endoplasmic Reticulum.Int J Mol Sci. 2022 Mar 29;23(7):3773. doi: 10.3390/ijms23073773. Int J Mol Sci. 2022. PMID: 35409131 Free PMC article. Review.
References
-
- Albanèse V., Yam A. Y., Baughman J., Parnot C., Frydman J. Systems analyses reveal two chaperone networks with distinct functions in eukaryotic cells. Cell. 2006;124:75–88. - PubMed
-
- Arnold C. E., Wittrup K. D. The stress response to loss of signal recognition particle function in Saccharomyces cerevisiae. J. Biol. Chem. 1994;269:30412–30418. - PubMed
-
- Bacher G., Lütcke H., Jungnickel B., Rapoport T. A., Dobberstein B. Regulation by the ribosome of the GTPase of the signal recognition particle during protein targeting. Nature. 1996;381:248–251. - PubMed
-
- Beckmann R., Spahn C.M.T., Eswar N., Helmers J., Penczek P. A., Sall A., Frank J., Blobel G. Architecture of the protein-conducting channel associated with the translating 80S ribosome. Cell. 2001;107:361–372. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

