Inter-kingdom conservation of mechanism of nonsense-mediated mRNA decay
- PMID: 18451801
- PMCID: PMC2426726
- DOI: 10.1038/emboj.2008.88
Inter-kingdom conservation of mechanism of nonsense-mediated mRNA decay
Abstract
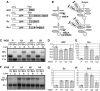
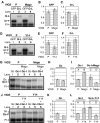
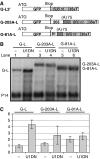
Nonsense-mediated mRNA decay (NMD) is a quality control system that degrades mRNAs containing premature termination codons. Although NMD is well characterized in yeast and mammals, plant NMD is poorly understood. We have undertaken the functional dissection of NMD pathways in plants. Using an approach that allows rapid identification of plant NMD trans factors, we demonstrated that two plant NMD pathways coexist, one eliminates mRNAs with long 3'UTRs, whereas a distinct pathway degrades mRNAs harbouring 3'UTR-located introns. We showed that UPF1, UPF2 and SMG-7 are involved in both plant NMD pathways, whereas Mago and Y14 are required only for intron-based NMD. The molecular mechanism of long 3'UTR-based plant NMD resembled yeast NMD, whereas the intron-based NMD was similar to mammalian NMD, suggesting that both pathways are evolutionarily conserved. Interestingly, the SMG-7 NMD component is targeted by NMD, suggesting that plant NMD is autoregulated. We propose that a complex, autoregulated NMD mechanism operated in stem eukaryotes, and that despite aspect of the mechanism being simplified in different lineages, feedback regulation was retained in all kingdoms.
Figures





Similar articles
-
Plant nonsense-mediated mRNA decay is controlled by different autoregulatory circuits and can be induced by an EJC-like complex.Nucleic Acids Res. 2013 Jul;41(13):6715-28. doi: 10.1093/nar/gkt366. Epub 2013 May 10. Nucleic Acids Res. 2013. PMID: 23666629 Free PMC article.
-
Both introns and long 3'-UTRs operate as cis-acting elements to trigger nonsense-mediated decay in plants.Nucleic Acids Res. 2006;34(21):6147-57. doi: 10.1093/nar/gkl737. Epub 2006 Nov 6. Nucleic Acids Res. 2006. PMID: 17088291 Free PMC article.
-
Control of mRNA Stability in Fungi by NMD, EJC and CBC Factors Through 3'UTR Introns.Genetics. 2015 Aug;200(4):1133-48. doi: 10.1534/genetics.115.176743. Epub 2015 Jun 4. Genetics. 2015. PMID: 26048019 Free PMC article.
-
Cutting the nonsense: the degradation of PTC-containing mRNAs.Biochem Soc Trans. 2010 Dec;38(6):1615-20. doi: 10.1042/BST0381615. Biochem Soc Trans. 2010. PMID: 21118136 Review.
-
Execution of nonsense-mediated mRNA decay: what defines a substrate?Curr Opin Cell Biol. 2009 Jun;21(3):394-402. doi: 10.1016/j.ceb.2009.02.007. Epub 2009 Apr 7. Curr Opin Cell Biol. 2009. PMID: 19359157 Review.
Cited by
-
Mechanism of cytoplasmic mRNA translation.Arabidopsis Book. 2015 Apr 24;13:e0176. doi: 10.1199/tab.0176. eCollection 2015. Arabidopsis Book. 2015. PMID: 26019692 Free PMC article.
-
Beyond Transcription: Fine-Tuning of Circadian Timekeeping by Post-Transcriptional Regulation.Genes (Basel). 2018 Dec 10;9(12):616. doi: 10.3390/genes9120616. Genes (Basel). 2018. PMID: 30544736 Free PMC article. Review.
-
The exon junction complex differentially marks spliced junctions.Nat Struct Mol Biol. 2010 Oct;17(10):1269-71. doi: 10.1038/nsmb.1890. Epub 2010 Sep 5. Nat Struct Mol Biol. 2010. PMID: 20818392
-
Noisy splicing, more than expression regulation, explains why some exons are subject to nonsense-mediated mRNA decay.BMC Biol. 2009 May 14;7:23. doi: 10.1186/1741-7007-7-23. BMC Biol. 2009. PMID: 19442261 Free PMC article.
-
Regulation of natural mRNAs by the nonsense-mediated mRNA decay pathway.Eukaryot Cell. 2014 Sep;13(9):1126-35. doi: 10.1128/EC.00090-14. Epub 2014 Jul 18. Eukaryot Cell. 2014. PMID: 25038084 Free PMC article. Review.
References
-
- Amrani N, Ganesan R, Kervestin S, Mangus DA, Ghosh S, Jacobson A (2004) A faux 3′-UTR promotes aberrant termination and triggers nonsense-mediated mRNA decay. Nature 432: 112–118 - PubMed
-
- Arciga-Reyes L, Wootton L, Kieffer M, Davies B (2006) UPF1 is required for nonsense-mediated mRNA decay (NMD) and RNAi in Arabidopsis. Plant J 47: 480–489 - PubMed
-
- Bono F, Ebert J, Lorentzen E, Conti E (2006) The crystal structure of the exon junction complex reveals how it maintains a stable grip on mRNA. Cell 126: 713–725 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

