Thioredoxin A active-site mutants form mixed disulfide dimers that resemble enzyme-substrate reaction intermediates
- PMID: 18455736
- PMCID: PMC2896474
- DOI: 10.1016/j.jmb.2008.03.077
Thioredoxin A active-site mutants form mixed disulfide dimers that resemble enzyme-substrate reaction intermediates
Abstract
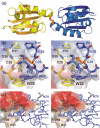
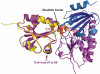
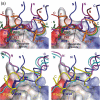
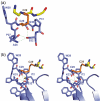
Thioredoxin functions in nearly all organisms as the major thiol-disulfide oxidoreductase within the cytosol. Its prime purpose is to maintain cysteine-containing proteins in the reduced state by converting intramolecular disulfide bonds into dithiols in a disulfide exchange reaction. Thioredoxin has been reported to contribute to a wide variety of physiological functions by interacting with specific sets of substrates in different cell types. To investigate the function of the essential thioredoxin A (TrxA) in the low-GC Gram-positive bacterium Bacillus subtilis, we purified wild-type TrxA and three mutant TrxA proteins that lack either one or both of the two cysteine residues in the CxxC active site. The pure proteins were used for substrate-binding studies known as "mixed disulfide fishing" in which covalent disulfide-bonded reaction intermediates can be visualized. An unprecedented finding is that both active-site cysteine residues can form mixed disulfides with substrate proteins when the other active-site cysteine is absent, but only the N-terminal active-site cysteine forms stable interactions. A second novelty is that both single-cysteine mutant TrxA proteins form stable homodimers due to thiol oxidation of the remaining active-site cysteine residue. To investigate whether these dimers resemble mixed enzyme-substrate disulfides, the structure of the most abundant dimer, C32S, was characterized by X-ray crystallography. This yielded a high-resolution (1.5A) X-ray crystallographic structure of a thioredoxin homodimer from a low-GC Gram-positive bacterium. The C32S TrxA dimer can be regarded as a mixed disulfide reaction intermediate of thioredoxin, which reveals the diversity of thioredoxin/substrate-binding modes.
Figures


Similar articles
-
The crystal structure of TrxA(CACA): Insights into the formation of a [2Fe-2S] iron-sulfur cluster in an Escherichia coli thioredoxin mutant.Protein Sci. 2005 Jul;14(7):1863-9. doi: 10.1110/ps.051464705. Protein Sci. 2005. PMID: 15987909 Free PMC article.
-
Mixed disulfide intermediates during the reduction of disulfides by Escherichia coli thioredoxin.Biochemistry. 1995 Sep 19;34(37):11807-13. doi: 10.1021/bi00037a019. Biochemistry. 1995. PMID: 7547914
-
Conformational fluctuations coupled to the thiol-disulfide transfer between thioredoxin and arsenate reductase in Bacillus subtilis.J Biol Chem. 2007 Apr 13;282(15):11078-83. doi: 10.1074/jbc.M700970200. Epub 2007 Feb 15. J Biol Chem. 2007. PMID: 17303556
-
Structural and mechanistic aspects of S-S bonds in the thioredoxin-like family of proteins.Biol Chem. 2019 Apr 24;400(5):575-587. doi: 10.1515/hsz-2018-0319. Biol Chem. 2019. PMID: 30367780 Review.
-
Protein disulfides and protein disulfide oxidoreductases in hyperthermophiles.FEBS J. 2006 Sep;273(18):4170-85. doi: 10.1111/j.1742-4658.2006.05421.x. Epub 2006 Aug 23. FEBS J. 2006. PMID: 16930136 Review.
Cited by
-
The Thioredoxin System of Mammalian Cells and Its Modulators.Biomedicines. 2022 Jul 21;10(7):1757. doi: 10.3390/biomedicines10071757. Biomedicines. 2022. PMID: 35885063 Free PMC article. Review.
-
Thioredoxin Txnl1/TRP32 is a redox-active cofactor of the 26 S proteasome.J Biol Chem. 2009 May 29;284(22):15246-54. doi: 10.1074/jbc.M900016200. Epub 2009 Apr 6. J Biol Chem. 2009. PMID: 19349277 Free PMC article.
-
Inhibitory Peptide of Soluble Guanylyl Cyclase/Trx1 Interface Blunts the Dual Redox Signaling Functions of the Complex.Antioxidants (Basel). 2023 Apr 10;12(4):906. doi: 10.3390/antiox12040906. Antioxidants (Basel). 2023. PMID: 37107281 Free PMC article.
-
Bacillus subtilis YtpP and Thioredoxin A Are New Players in the Coenzyme-A-Mediated Defense Mechanism against Cellular Stress.Antioxidants (Basel). 2023 Apr 15;12(4):938. doi: 10.3390/antiox12040938. Antioxidants (Basel). 2023. PMID: 37107313 Free PMC article.
-
Molecular dissection of the Erv41-Erv46 retrograde receptor reveals a conserved cysteine-rich region in Erv46 required for retrieval activity.Mol Biol Cell. 2020 Feb 1;31(3):209-220. doi: 10.1091/mbc.E19-08-0484. Epub 2019 Dec 11. Mol Biol Cell. 2020. PMID: 31825724 Free PMC article.
References
-
- Arner E.S., Holmgren A. Physiological functions of thioredoxin and thioredoxin reductase. Eur. J. Biochem. 2000;267:6102–6109. - PubMed
-
- Laurent T.C., Moore E.C., Richard P. Enzymatic synthesis of deoxyribonucleotides: IV. Isolation and characterization of thioredoxin, the hydrogen donor from Escherichia coli B. J. Biol. Chem. 1964;239:3436–3444. - PubMed
-
- Powis G., Mustacich D., Coon A. The role of the redox protein thioredoxin in cell growth and cancer. Free Radic. Biol. Med. 2000;29:312–322. - PubMed
-
- Schurmann P. Redox signaling in the chloroplast: the ferredoxin/thioredoxin system. Antioxid. Redox Signal. 2003;5:69–78. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

