SRP keeps polypeptides translocation-competent by slowing translation to match limiting ER-targeting sites
- PMID: 18455985
- PMCID: PMC2430734
- DOI: 10.1016/j.cell.2008.02.049
SRP keeps polypeptides translocation-competent by slowing translation to match limiting ER-targeting sites
Abstract
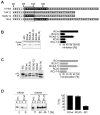
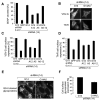
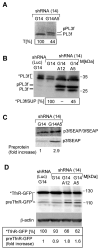
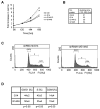
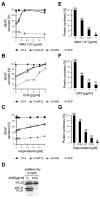
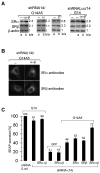
SRP is essential for targeting nascent chains to the endoplasmic reticulum, and it delays nascent chain elongation in cell-free translation systems. However, the significance of this function has remained unclear. We show that efficient protein translocation into the ER is incompatible with normal cellular translation rates due to rate-limiting concentrations of SRP receptor (SR). We complemented mammalian cells depleted of SRP14 by expressing mutant versions of the protein lacking the elongation arrest function. The absence of a delay caused inefficient targeting of preproteins leading to defects in secretion, depletion of proteins in the endogenous membranes, and reduced cell growth. The detrimental effects were reversed by either reducing the cellular protein synthesis rate or increasing SR expression. SRP therefore ensures that nascent chains remain translocation competent during the targeting time window dictated by SR. Since SRP-signal sequence affinities vary, the delay may also regulate which proteins are preferentially targeted.
Figures
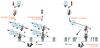
References
-
- Alder NN, Johnson AE. Cotranslational membrane protein biogenesis at the endoplasmic reticulum. J Biol Chem. 2004;279:22787–22790. - PubMed
-
- Blau M, Mullapudi S, Becker T, Dudek J, Zimmermann R, Penczek PA, Beckmann R. ERj1p uses a universal ribosomal adaptor site to coordinate the 80S ribosome at the membrane. Nat Struct Mol Biol. 2005;12:1015–1016. - PubMed
-
- Bordeleau ME, Mori A, Oberer M, Lindqvist L, Chard LS, Higa T, Belsham GJ, Wagner G, Tanaka J, Pelletier J. Functional characterization of IRESes by an inhibitor of the RNA helicase eIF4A. Nat Chem Biol. 2006;2:213–220. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

