The stream of precursors that colonizes the thymus proceeds selectively through the early T lineage precursor stage of T cell development
- PMID: 18458114
- PMCID: PMC2373849
- DOI: 10.1084/jem.20072168
The stream of precursors that colonizes the thymus proceeds selectively through the early T lineage precursor stage of T cell development
Abstract
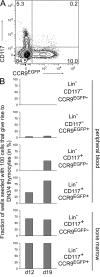
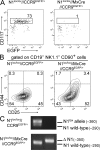
T cell development in the thymus depends on continuous colonization by hematopoietic precursors. Several distinct T cell precursors have been identified, but whether one or several independent precursor cell types maintain thymopoiesis is unclear. We have used thymus transplantation and an inducible lineage-tracing system to identify the intrathymic precursor cells among previously described thymus-homing progenitors that give rise to the T cell lineage in the thymus. Extrathymic precursors were not investigated in these studies. Both approaches show that the stream of T cell lineage precursor cells, when entering the thymus, selectively passes through the early T lineage precursor (ETP) stage. Immigrating precursor cells do not exhibit characteristics of double-negative (DN) 1c, DN1d, or DN1e stages, or of populations containing the common lymphoid precursor 2 (CLP-2) or the thymic equivalent of circulating T cell progenitors (CTPs). It remains possible that an unknown hematopoietic precursor cell or previously described extrathymic precursors with a CLP, CLP-2, or CTP phenotype feed into T cell development by circumventing known intrathymic T cell lineage progenitor cells. However, it is clear that of the known intrathymic precursors, only the ETP population contributes significant numbers of T lineage precursors to T cell development.
Figures




References
-
- Bhandoola, A., H. von Boehmer, H.T. Petrie, J.C. Zuniga-Pflucker, V.P. Zediak, I. Maillard, A. Bhandoola, B.A. Schwarz, A. Sambandam, I. Maillard, et al. 2007. Commitment and developmental potential of extrathymic and intrathymic T cell precursors: plenty to choose from. Immunity. 26:678–689. - PubMed
-
- Boehm, T., and C.C. Bleul. 2006. Thymus-homing precursors and the thymic microenvironment. Trends Immunol. 27:477–484. - PubMed
-
- Wu, L., R. Scollay, M. Egerton, M. Pearse, G.J. Spangrude, and K. Shortman. 1991. CD4 expressed on earliest T-lineage precursor cells in the adult murine thymus. Nature. 349:71–74. - PubMed
-
- Allman, D., A. Sambandam, S. Kim, J.P. Miller, A. Pagan, D. Well, A. Meraz, and A. Bhandoola. 2003. Thymopoiesis independent of common lymphoid progenitors. Nat. Immunol. 4:168–174. - PubMed
-
- Martin, C.H., I. Aifantis, M.L. Scimone, U.H. von Andrian, B. Reizis, H. von Boehmer, and F. Gounari. 2003. Efficient thymic immigration of B220(+) lymphoid-restricted bone marrow cells with T precursor potential. Nat. Immunol. 4:866–873. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

