De-orphaning the structural proteome through reciprocal comparison of evolutionarily important structural features
- PMID: 18461181
- PMCID: PMC2362850
- DOI: 10.1371/journal.pone.0002136
De-orphaning the structural proteome through reciprocal comparison of evolutionarily important structural features
Abstract
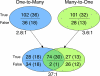
Function prediction frequently relies on comparing genes or gene products to search for relevant similarities. Because the number of protein structures with unknown function is mushrooming, however, we asked here whether such comparisons could be improved by focusing narrowly on the key functional features of protein structures, as defined by the Evolutionary Trace (ET). Therefore a series of algorithms was built to (a) extract local motifs (3D templates) from protein structures based on ET ranking of residue importance; (b) to assess their geometric and evolutionary similarity to other structures; and (c) to transfer enzyme annotation whenever a plurality was reached across matches. Whereas a prototype had only been 80% accurate and was not scalable, here a speedy new matching algorithm enabled large-scale searches for reciprocal matches and thus raised annotation specificity to 100% in both positive and negative controls of 49 enzymes and 50 non-enzymes, respectively-in one case even identifying an annotation error-while maintaining sensitivity ( approximately 60%). Critically, this Evolutionary Trace Annotation (ETA) pipeline requires no prior knowledge of functional mechanisms. It could thus be applied in a large-scale retrospective study of 1218 structural genomics enzymes and reached 92% accuracy. Likewise, it was applied to all 2935 unannotated structural genomics proteins and predicted enzymatic functions in 320 cases: 258 on first pass and 62 more on second pass. Controls and initial analyses suggest that these predictions are reliable. Thus the large-scale evolutionary integration of sequence-structure-function data, here through reciprocal identification of local, functionally important structural features, may contribute significantly to de-orphaning the structural proteome.
Conflict of interest statement
Figures


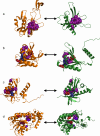
References
-
- Chandonia JM, Brenner SE. The impact of structural genomics: expectations and outcomes. Science. 2006;311:347–351. - PubMed
-
- Brenner SE. A tour of structural genomics. Nat Rev Genet. 2001;2:801–809. - PubMed
-
- Burley SK. An overview of structural genomics. Nat Struct Biol. 2000;7(Suppl):932–934. - PubMed
-
- Leulliot N, Tresaugues L, Bremang M, Sorel I, Ulryck N, et al. High-throughput crystal-optimization strategies in the South Paris Yeast Structural Genomics Project: one size fits all? Acta Crystallogr D Biol Crystallogr. 2005;61:664–670. - PubMed
-
- Baker D, Sali A. Protein structure prediction and structural genomics. Science. 2001;294:93–96. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

