Accurate localization of the integration sites of two genomic islands at single-nucleotide resolution in the genome of Bacillus cereus ATCC 10987
- PMID: 18464912
- PMCID: PMC2359905
- DOI: 10.1155/2008/451930
Accurate localization of the integration sites of two genomic islands at single-nucleotide resolution in the genome of Bacillus cereus ATCC 10987
Abstract
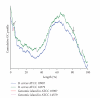
We have identified two genomic islands, that is, BCEGI-1 and BCEGI-2, in the genome of Bacillus cereus ATCC 10987, based on comparative analysis with Bacillus cereus ATCC 14579. Furthermore, by using the cumulative GC profile and performing homology searches between the two genomes, the integration sites of the two genomic islands were determined at single-nucleotide resolution. BCEGI-1 is integrated between 159705 bp and 198000 bp, whereas BCEGI-2 is integrated between the end of ORF BCE4594 and the start of the intergenic sequence immediately following BCE4626, that is, from 4256803 bp to 4285534 bp. BCEGI-1 harbors two bacterial Tn7 transposons, which have two sets of genes encoding TnsA, B, C, and D. It is generally believed that unlike the TnsABC+E pathway, the TnsABC+D pathway would only promote vertical transmission to daughter cells. The evidence presented in this paper, however, suggests a role of the TnsABC+D pathway in the horizontal transfer of some genomic islands.
Figures
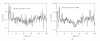



Similar articles
-
Identification of genomic islands in the genome of Bacillus cereus by comparative analysis with Bacillus anthracis.Physiol Genomics. 2003 Dec 16;16(1):19-23. doi: 10.1152/physiolgenomics.00170.2003. Physiol Genomics. 2003. PMID: 14600214
-
The bcr1 DNA repeat element is specific to the Bacillus cereus group and exhibits mobile element characteristics.J Bacteriol. 2004 Nov;186(22):7714-25. doi: 10.1128/JB.186.22.7714-7725.2004. J Bacteriol. 2004. PMID: 15516586 Free PMC article.
-
Novel Effective Bacillus cereus Group Species "Bacillus clarus" Is Represented by Antibiotic-Producing Strain ATCC 21929 Isolated from Soil.mSphere. 2020 Nov 4;5(6):e00882-20. doi: 10.1128/mSphere.00882-20. mSphere. 2020. PMID: 33148822 Free PMC article.
-
Unusual group II introns in bacteria of the Bacillus cereus group.J Bacteriol. 2005 Aug;187(15):5437-51. doi: 10.1128/JB.187.15.5437-5451.2005. J Bacteriol. 2005. PMID: 16030238 Free PMC article.
-
Heteromeric transposase elements: generators of genomic islands across diverse bacteria.Mol Microbiol. 2014 Sep;93(6):1084-92. doi: 10.1111/mmi.12740. Epub 2014 Aug 19. Mol Microbiol. 2014. PMID: 25091064 Review.
Cited by
-
Bacillus cytotoxicus Genomics: Chromosomal Diversity and Plasmidome Versatility.Front Microbiol. 2021 Dec 9;12:789929. doi: 10.3389/fmicb.2021.789929. eCollection 2021. Front Microbiol. 2021. PMID: 34992589 Free PMC article.
-
A novel non prophage(-like) gene-intervening element within gerE that is reconstituted during sporulation in Bacillus cereus ATCC10987.Sci Rep. 2017 Sep 12;7(1):11426. doi: 10.1038/s41598-017-11796-8. Sci Rep. 2017. PMID: 28900282 Free PMC article.
-
Prediction of genomic islands in three bacterial pathogens of pneumonia.Int J Mol Sci. 2012;13(3):3134-3144. doi: 10.3390/ijms13033134. Epub 2012 Mar 7. Int J Mol Sci. 2012. PMID: 22489145 Free PMC article.
-
Zisland Explorer: detect genomic islands by combining homogeneity and heterogeneity properties.Brief Bioinform. 2017 May 1;18(3):357-366. doi: 10.1093/bib/bbw019. Brief Bioinform. 2017. PMID: 26992782 Free PMC article.
-
Identification of Horizontally-transferred Genomic Islands and Genome Segmentation Points by Using the GC Profile Method.Curr Genomics. 2014 Apr;15(2):113-21. doi: 10.2174/1389202915999140328163125. Curr Genomics. 2014. PMID: 24822029 Free PMC article.
References
-
- Kotiranta A, Lounatmaa K, Haapasalo M. Epidemiology and pathogenesis of Bacillus cereus infections. Microbes and Infection. 2000;2(2):189–198. - PubMed
-
- Jensen GB, Hansen BM, Eilenberg J, Mahillon J. The hidden lifestyles of Bacillus cereus and relatives. Environmental Microbiology. 2003;5(8):631–640. - PubMed
-
- Ivanova N, Sorokin A, Anderson I, et al. Genome sequence of Bacillus cereus and comparative analysis with Bacillus anthracis. Nature. 2003;423(6935):87–91. - PubMed
-
- Hacker J, Kaper JB. Pathogenicity islands and the evolution of microbes. Annual Review of Microbiology. 2000;54:641–679. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

