Sequence diversity in the A domain of Staphylococcus aureus fibronectin-binding protein A
- PMID: 18466610
- PMCID: PMC2390562
- DOI: 10.1186/1471-2180-8-74
Sequence diversity in the A domain of Staphylococcus aureus fibronectin-binding protein A
Abstract
Background: Fibronectin-binding protein A (FnBPA) mediates adhesion of Staphylococcus aureus to fibronectin, fibrinogen and elastin. We previously reported that S. aureus strain P1 encodes an FnBPA protein where the fibrinogen/elastin-binding domain (A domain) is substantially divergent in amino acid sequence from the archetypal FnBPA of S. aureus NCTC8325, and that these variations created differences in antigenicity. In this study strains from multilocus sequence types (MLST) that spanned the genetic diversity of S.aureus were examined to determine the extent of FnBPA A domain variation within the S. aureus population and its effect on ligand binding and immuno-crossreactivity.
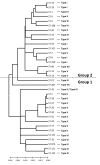
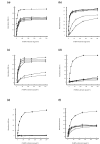
Results: Seven different isotype forms (I - VII) of the FnBPA A domain were identified which were between 66 to 76% identical in amino acid sequence in any pair-wise alignment. The fnbA allelic variants in strains of different multilocus sequence type were identified by DNA hybridization using probes specific for sequences encoding the highly divergent N3 sub-domain of different isotypes. Several isotypes were not restricted to specific clones or clonal complexes but were more widely distributed. It is highly likely that certain fnbA genes have been transferred horizontally. Residues lining the putative ligand-binding trench were conserved, which is consistent with the ability of each A domain isotype to bind immobilized fibrinogen and elastin by the dock-latch-lock mechanism. Variant amino acid residues were mapped on a three-dimensional model of the FnBPA A domain and were predicted to be surface-exposed. Polyclonal antibodies raised against the recombinant isotype I A domain bound that protein with a 4 - 7 fold higher apparent affinity compared to the A domains of isotypes II - VII, while some monoclonal antibodies generated against the isotype I A domain showed reduced or no binding to the other isotypes.
Conclusion: The FnBPA A domain occurs in at least 7 different isotypes which differ antigenically and exhibit limited immuno-crossreactivity, yet retain their ligand-binding functions. Antigenic variation of the FnBPA A domain may aid S. aureus to evade the host's immune responses. These findings have implications for the development of vaccines or immunotherapeutics that target FnBPA.
Figures


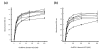
Similar articles
-
Fibronectin-binding protein B variation in Staphylococcus aureus.BMC Microbiol. 2010 Jun 1;10:160. doi: 10.1186/1471-2180-10-160. BMC Microbiol. 2010. PMID: 20515471 Free PMC article.
-
Variation and association of fibronectin-binding protein genes fnbA and fnbB in Staphylococcus aureus Japanese isolates.Microbiol Immunol. 2016 May;60(5):312-25. doi: 10.1111/1348-0421.12377. Microbiol Immunol. 2016. PMID: 26990092
-
Fibrinogen and elastin bind to the same region within the A domain of fibronectin binding protein A, an MSCRAMM of Staphylococcus aureus.Mol Microbiol. 2007 Feb;63(3):711-23. doi: 10.1111/j.1365-2958.2006.05552.x. Mol Microbiol. 2007. PMID: 17302800
-
The remarkably multifunctional fibronectin binding proteins of Staphylococcus aureus.Eur J Clin Microbiol Infect Dis. 2016 Dec;35(12):1923-1931. doi: 10.1007/s10096-016-2763-0. Epub 2016 Sep 7. Eur J Clin Microbiol Infect Dis. 2016. PMID: 27604831 Review.
-
The Multivalent Role of Fibronectin-Binding Proteins A and B (FnBPA and FnBPB) of Staphylococcus aureus in Host Infections.Front Microbiol. 2020 Aug 26;11:2054. doi: 10.3389/fmicb.2020.02054. eCollection 2020. Front Microbiol. 2020. PMID: 32983039 Free PMC article. Review.
Cited by
-
Whole-Genome Sequencing and Machine Learning Analysis of Staphylococcus aureus from Multiple Heterogeneous Sources in China Reveals Common Genetic Traits of Antimicrobial Resistance.mSystems. 2021 Jun 29;6(3):e0118520. doi: 10.1128/mSystems.01185-20. Epub 2021 Jun 8. mSystems. 2021. PMID: 34100643 Free PMC article.
-
Molecular characterisation of virulence genes in bacterial pathogens from daycare centres in Ile-Ife, Nigeria: implications for infection control.BMC Infect Dis. 2024 Oct 23;24(1):1196. doi: 10.1186/s12879-024-10095-8. BMC Infect Dis. 2024. PMID: 39443869 Free PMC article.
-
Fibronectin-binding protein B variation in Staphylococcus aureus.BMC Microbiol. 2010 Jun 1;10:160. doi: 10.1186/1471-2180-10-160. BMC Microbiol. 2010. PMID: 20515471 Free PMC article.
-
Fibronectin-binding protein B (FnBPB) from Staphylococcus aureus protects against the antimicrobial activity of histones.J Biol Chem. 2019 Mar 8;294(10):3588-3602. doi: 10.1074/jbc.RA118.005707. Epub 2019 Jan 8. J Biol Chem. 2019. PMID: 30622139 Free PMC article.
-
Combining use of a panel of ssDNA aptamers in the detection of Staphylococcus aureus.Nucleic Acids Res. 2009 Aug;37(14):4621-8. doi: 10.1093/nar/gkp489. Epub 2009 Jun 4. Nucleic Acids Res. 2009. PMID: 19498077 Free PMC article.
References
-
- Signas C, Raucci G, Jonsson K, Lindgren PE, Anantharamaiah GM, Hook M, Lindberg M. Nucleotide sequence of the gene for a fibronectin-binding protein from Staphylococcus aureus: use of this peptide sequence in the synthesis of biologically active peptides. Proc Natl Acad Sci U S A. 1989;86:699–703. doi: 10.1073/pnas.86.2.699. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

